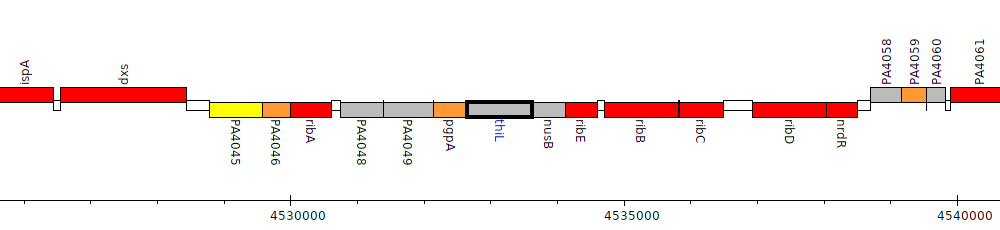
Pseudomonas aeruginosa PAO1, PA4051 (thiL)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0009030 | thiamine-phosphate kinase activity | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
9188462 | Reviewed by curator |
| Biological Process | GO:0051188 | obsolete cofactor biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0009030 | thiamine-phosphate kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_02128
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009228 | thiamine biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:MF_02128
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCyc | THISYN-PWY | superpathway of thiamin diphosphate biosynthesis I | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | PWY-6894 | thiamin diphosphate biosynthesis I (E. coli) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Thiamine metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00730 | Thiamine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00586 | AIR synthase related protein, N-terminal domain | IPR016188 | PurM-like, N-terminal domain | 29 | 138 | 3.5E-25 |
| CDD | cd02194 | ThiL | IPR006283 | Thiamine-monophosphate kinase-like | 2 | 292 | 9.14679E-109 |
| PANTHER | PTHR30270 | THIAMINE-MONOPHOSPHATE KINASE | IPR006283 | Thiamine-monophosphate kinase-like | 2 | 319 | 1.5E-75 |
| Gene3D | G3DSA:3.30.1330.10 | - | IPR036921 | PurM-like, N-terminal domain superfamily | 1 | 144 | 2.9E-48 |
| Hamap | MF_02128 | Thiamine-monophosphate kinase [thiL]. | IPR006283 | Thiamine-monophosphate kinase-like | 1 | 317 | 27.013096 |
| SUPERFAMILY | SSF56042 | PurM C-terminal domain-like | IPR036676 | PurM-like, C-terminal domain superfamily | 143 | 303 | 6.49E-35 |
| Gene3D | G3DSA:3.90.650.10 | - | IPR036676 | PurM-like, C-terminal domain superfamily | 145 | 317 | 4.8E-45 |
| PIRSF | PIRSF005303 | Thiam_monoph_kin | IPR006283 | Thiamine-monophosphate kinase-like | 1 | 317 | 5.2E-78 |
| NCBIfam | TIGR01379 | JCVI: thiamine-phosphate kinase | IPR006283 | Thiamine-monophosphate kinase-like | 2 | 317 | 1.8E-94 |
| SUPERFAMILY | SSF55326 | PurM N-terminal domain-like | IPR036921 | PurM-like, N-terminal domain superfamily | 1 | 138 | 2.09E-34 |
| Pfam | PF02769 | AIR synthase related protein, C-terminal domain | IPR010918 | PurM-like, C-terminal domain | 150 | 297 | 3.6E-6 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.