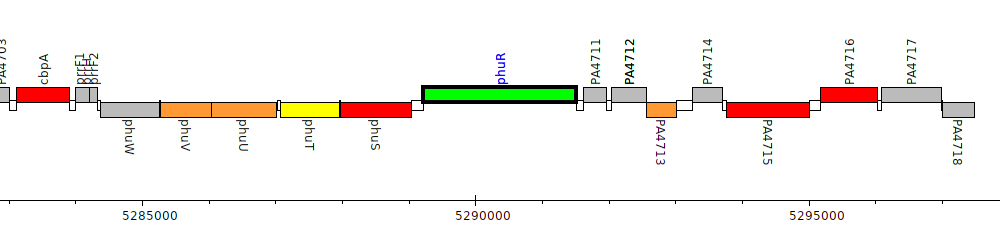
Pseudomonas aeruginosa PAO1, PA4710 (phuR)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0022857 | transmembrane transporter activity | |
|||
| Cellular Component | GO:0019867 | outer membrane | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
10658665 | Reviewed by curator |
| Biological Process | GO:0015886 | heme transport | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
10658665 | Reviewed by curator |
| Molecular Function | GO:0015232 | heme transmembrane transporter activity | Inferred from Mutant Phenotype | ECO:0000315 mutant phenotype evidence used in manual assertion |
10658665 | Reviewed by curator |
| Molecular Function | GO:0015232 | heme transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01785
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01786
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0022857 | transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01786
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0015886 | heme transport |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01785
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0019867 | outer membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01785
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| NCBIfam | TIGR01786 | JCVI: TonB-dependent hemoglobin/transferrin/lactoferrin family receptor | IPR010949 | TonB-dependent haemoglobin/transferrin/lactoferrin receptor | 34 | 761 | 0.0 |
| PANTHER | PTHR30069 | TONB-DEPENDENT OUTER MEMBRANE RECEPTOR | IPR039426 | TonB-dependent receptor-like | 14 | 762 | 2.1E-122 |
| Gene3D | G3DSA:2.40.170.20 | - | IPR036942 | TonB-dependent receptor-like, beta-barrel domain superfamily | 171 | 763 | 2.7E-129 |
| SUPERFAMILY | SSF56935 | Porins | - | - | 35 | 760 | 2.29E-122 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 397 | 420 | - |
| CDD | cd01347 | ligand_gated_channel | - | - | 51 | 763 | 7.0357E-107 |
| Pfam | PF07715 | TonB-dependent Receptor Plug Domain | IPR012910 | TonB-dependent receptor, plug domain | 46 | 155 | 1.2E-27 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 212 | 233 | - |
| Gene3D | G3DSA:2.170.130.10 | - | IPR037066 | TonB-dependent receptor, plug domain superfamily | 47 | 170 | 1.4E-34 |
| NCBIfam | TIGR01785 | JCVI: TonB-dependent heme/hemoglobin receptor family protein | IPR011276 | TonB-dependent haem/haemoglobin receptor | 33 | 760 | 5.3E-126 |
| Pfam | PF00593 | TonB dependent receptor | IPR000531 | TonB-dependent receptor-like, beta-barrel | 293 | 761 | 4.1E-65 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.