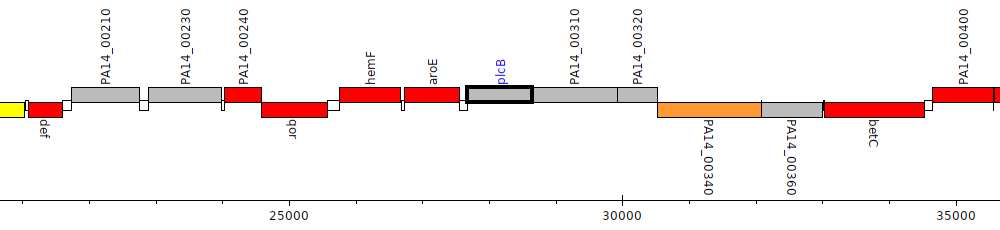
Pseudomonas aeruginosa UCBPP-PA14, PA14_00300 (plcB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006935 | chemotaxis |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA0026
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
15306013 | |
| Molecular Function | GO:0004629 | phospholipase C activity |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA0026
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
15306013 | |
| Molecular Function | GO:0008270 | zinc ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd11009
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016788 | hydrolase activity, acting on ester bonds |
Inferred from Sequence Model
Term mapped from: InterPro:SSF48537
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004629 | phospholipase C activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd11009
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd11009 | Zn_dep_PLPC | IPR001531 | Zinc-dependent phospholipase C | 120 | 317 | 5.9609E-30 |
| Gene3D | G3DSA:1.10.575.10 | P1 Nuclease | IPR008947 | Phospholipase C/P1 nuclease domain superfamily | 145 | 325 | 7.9E-11 |
| SUPERFAMILY | SSF48537 | Phospholipase C/P1 nuclease | IPR008947 | Phospholipase C/P1 nuclease domain superfamily | 133 | 320 | 7.06E-34 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.