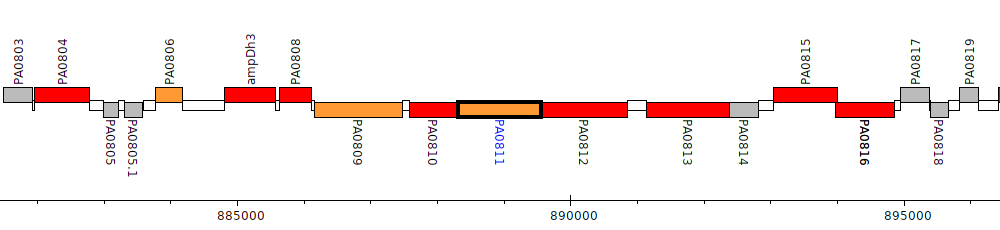
Pseudomonas aeruginosa PAO1, PA0811
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0022857 | transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF07690
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0055085 | transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:PF07690
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:1.20.1250.20 | MFS general substrate transporter like domains | IPR036259 | MFS transporter superfamily | 216 | 406 | 8.5E-13 |
| Gene3D | G3DSA:1.20.1250.20 | MFS general substrate transporter like domains | IPR036259 | MFS transporter superfamily | 7 | 199 | 5.3E-33 |
| SUPERFAMILY | SSF103473 | MFS general substrate transporter | IPR036259 | MFS transporter superfamily | 7 | 405 | 4.84E-61 |
| CDD | cd17475 | MFS_MT3072_like | - | - | 21 | 401 | 1.88938E-118 |
| Pfam | PF07690 | Major Facilitator Superfamily | IPR011701 | Major facilitator superfamily | 22 | 369 | 4.3E-48 |
| PANTHER | PTHR23527 | BLL3282 PROTEIN | - | - | 1 | 415 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.