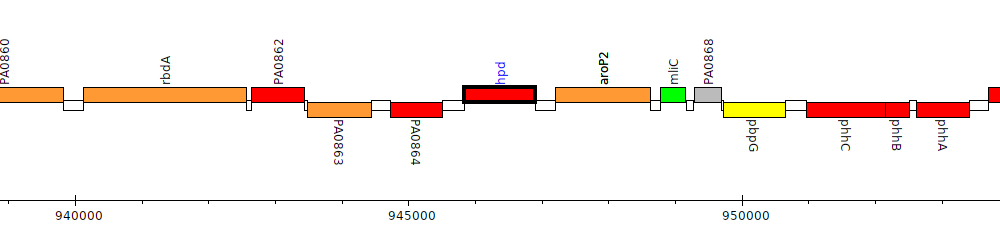
Pseudomonas aeruginosa PAO1, PA0865 (hpd)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006520 | cellular amino acid metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0016763 | transferase activity, transferring pentosyl groups | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Molecular Function | GO:0003868 | 4-hydroxyphenylpyruvate dioxygenase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01263
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016701 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01263
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009072 | aromatic amino acid family metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01263
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae00350 | Tyrosine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00130 | Ubiquinone and other terpenoid-quinone biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00360 | Phenylalanine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF14696 | Hydroxyphenylpyruvate dioxygenase, HPPD, N-terminal | - | - | 10 | 149 | 3.3E-52 |
| CDD | cd08342 | HPPD_N_like | IPR041736 | 4-hydroxyphenylpyruvate dioxygenase, N-terminal | 18 | 150 | 1.55075E-39 |
| PIRSF | PIRSF009283 | 4HPP_dioxygenase | IPR005956 | 4-hydroxyphenylpyruvate dioxygenase | 1 | 357 | 8.5E-121 |
| NCBIfam | TIGR01263 | JCVI: 4-hydroxyphenylpyruvate dioxygenase | IPR005956 | 4-hydroxyphenylpyruvate dioxygenase | 17 | 356 | 8.0E-124 |
| SUPERFAMILY | SSF54593 | Glyoxalase/Bleomycin resistance protein/Dihydroxybiphenyl dioxygenase | IPR029068 | Glyoxalase/Bleomycin resistance protein/Dihydroxybiphenyl dioxygenase | 8 | 344 | 3.29E-80 |
| FunFam | G3DSA:3.10.180.10:FF:000018 | 4-hydroxyphenylpyruvate dioxygenase | - | - | 8 | 160 | 9.3E-71 |
| CDD | cd07250 | HPPD_C_like | IPR041735 | 4-hydroxyphenylpyruvate dioxygenase, C-terminal | 163 | 349 | 3.09447E-87 |
| Gene3D | G3DSA:3.10.180.10 | - | IPR029068 | Glyoxalase/Bleomycin resistance protein/Dihydroxybiphenyl dioxygenase | 8 | 160 | 1.7E-47 |
| FunFam | G3DSA:3.10.180.10:FF:000007 | 4-hydroxyphenylpyruvate dioxygenase | - | - | 161 | 357 | 1.6E-110 |
| Gene3D | G3DSA:3.10.180.10 | - | IPR029068 | Glyoxalase/Bleomycin resistance protein/Dihydroxybiphenyl dioxygenase | 161 | 357 | 8.4E-72 |
| Pfam | PF00903 | Glyoxalase/Bleomycin resistance protein/Dioxygenase superfamily | IPR004360 | Glyoxalase/fosfomycin resistance/dioxygenase domain | 166 | 277 | 2.2E-16 |
| PANTHER | PTHR11959 | 4-HYDROXYPHENYLPYRUVATE DIOXYGENASE | IPR005956 | 4-hydroxyphenylpyruvate dioxygenase | 12 | 355 | 4.0E-75 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.