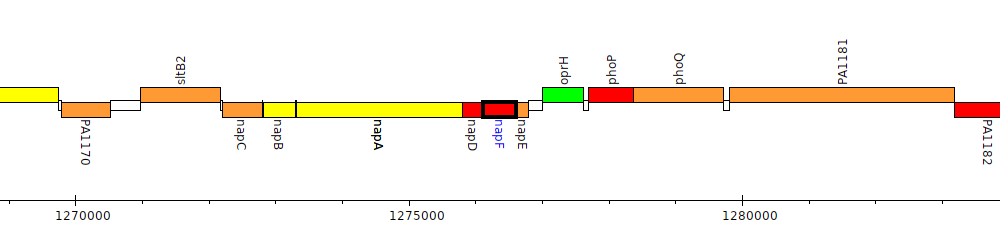
Pseudomonas aeruginosa PAO1, PA1176 (napF)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0009325 | nitrate reductase complex | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
15371424 | Reviewed by curator |
| Molecular Function | GO:0051539 | 4 iron, 4 sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd10564
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Nitrogen metabolism |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF12838 | 4Fe-4S dicluster domain | IPR017896 | 4Fe-4S ferredoxin-type, iron-sulphur binding domain | 31 | 76 | 1.0E-6 |
| CDD | cd10564 | NapF_like | IPR004496 | Ferredoxin-type protein NapF | 18 | 154 | 4.31732E-56 |
| Pfam | PF12838 | 4Fe-4S dicluster domain | IPR017896 | 4Fe-4S ferredoxin-type, iron-sulphur binding domain | 101 | 148 | 6.3E-9 |
| PANTHER | PTHR43687 | ADENYLYLSULFATE REDUCTASE, BETA SUBUNIT | - | - | 61 | 121 | 1.1E-10 |
| FunFam | G3DSA:3.30.70.20:FF:000025 | Ferredoxin-type protein NapF | - | - | 88 | 157 | 3.9E-33 |
| Gene3D | G3DSA:3.30.70.20 | - | - | - | 90 | 161 | 1.3E-9 |
| Gene3D | G3DSA:3.30.70.20 | - | - | - | 27 | 78 | 1.1E-8 |
| Hamap | MF_02201 | Ferredoxin-type protein NapF [napF]. | IPR004496 | Ferredoxin-type protein NapF | 21 | 155 | 21.950979 |
| NCBIfam | TIGR00402 | JCVI: ferredoxin-type protein NapF | IPR004496 | Ferredoxin-type protein NapF | 18 | 149 | 5.5E-31 |
| SUPERFAMILY | SSF54862 | 4Fe-4S ferredoxins | - | - | 28 | 151 | 9.77E-18 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.