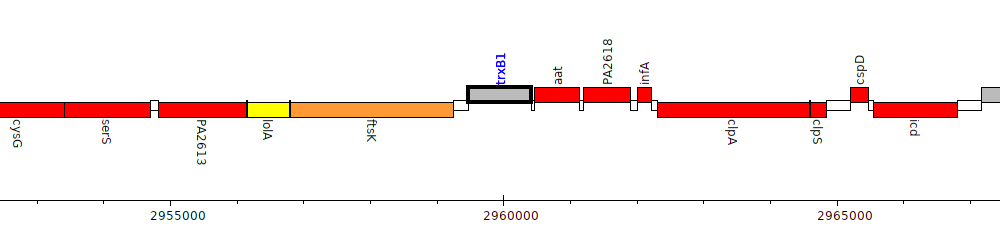
Pseudomonas aeruginosa PAO1, PA2616 (trxB1)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004356 | glutamate-ammonia ligase activity | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0006220 | pyrimidine nucleotide metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0009117 | nucleotide metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0019430 | removal of superoxide radicals |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01292
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01292
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF07992
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004791 | thioredoxin-disulfide reductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01292
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae00240 | Pyrimidine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00450 | Selenocompound metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Pyrimidine metabolism |
ECO:0000037
not_recorded |
|||
| PseudoCyc | THIOREDOX-PWY | thioredoxin pathway | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 107 | 115 | 1.1E-76 |
| Pfam | PF07992 | Pyridine nucleotide-disulphide oxidoreductase | IPR023753 | FAD/NAD(P)-binding domain | 8 | 300 | 4.4E-47 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 8 | 30 | 1.1E-76 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 9 | 28 | 1.5E-38 |
| NCBIfam | TIGR01292 | JCVI: thioredoxin-disulfide reductase | IPR005982 | Thioredoxin reductase | 8 | 313 | 1.2E-119 |
| SUPERFAMILY | SSF51905 | FAD/NAD(P)-binding domain | IPR036188 | FAD/NAD(P)-binding domain superfamily | 2 | 313 | 1.44E-55 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 276 | 294 | 1.1E-76 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 199 | 215 | 1.1E-76 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 9 | 313 | 0.0 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 266 | 288 | 1.5E-38 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 238 | 259 | 1.1E-76 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 62 | 72 | 1.1E-76 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 148 | 166 | 1.5E-38 |
| PANTHER | PTHR48105 | THIOREDOXIN REDUCTASE 1-RELATED-RELATED | - | - | 6 | 314 | 1.2E-100 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 106 | 124 | 1.5E-38 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 144 | 168 | 1.1E-76 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 236 | 252 | 1.5E-38 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 129 | 141 | 1.1E-76 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 118 | 244 | 0.0 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 41 | 56 | 1.1E-76 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.