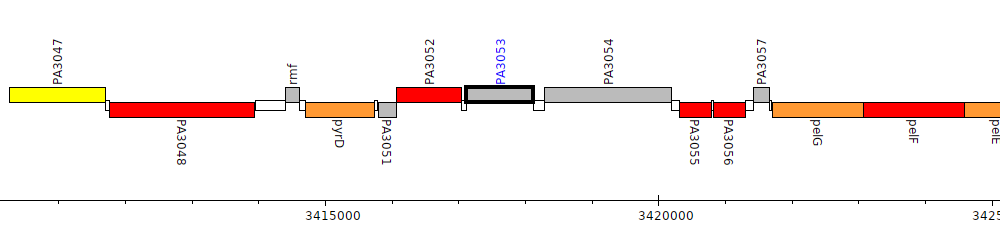
Pseudomonas aeruginosa PAO1, PA3053
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00412
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR43798 | MONOACYLGLYCEROL LIPASE | - | - | 61 | 332 | 1.7E-39 |
| SUPERFAMILY | SSF53474 | alpha/beta-Hydrolases | IPR029058 | Alpha/Beta hydrolase fold | 47 | 331 | 4.96E-62 |
| Gene3D | G3DSA:3.40.50.1820 | alpha/beta hydrolase | IPR029058 | Alpha/Beta hydrolase fold | 21 | 334 | 1.4E-115 |
| PRINTS | PR00412 | Epoxide hydrolase signature | IPR000639 | Epoxide hydrolase-like | 152 | 165 | 1.8E-6 |
| PRINTS | PR00412 | Epoxide hydrolase signature | IPR000639 | Epoxide hydrolase-like | 138 | 151 | 1.8E-6 |
| PRINTS | PR00111 | Alpha/beta hydrolase fold signature | IPR000073 | Alpha/beta hydrolase fold-1 | 152 | 165 | 1.5E-11 |
| PRINTS | PR00412 | Epoxide hydrolase signature | IPR000639 | Epoxide hydrolase-like | 73 | 91 | 1.8E-6 |
| PRINTS | PR00412 | Epoxide hydrolase signature | IPR000639 | Epoxide hydrolase-like | 93 | 108 | 1.8E-6 |
| PRINTS | PR00111 | Alpha/beta hydrolase fold signature | IPR000073 | Alpha/beta hydrolase fold-1 | 93 | 108 | 1.5E-11 |
| Pfam | PF00561 | alpha/beta hydrolase fold | IPR000073 | Alpha/beta hydrolase fold-1 | 68 | 319 | 1.3E-23 |
| PRINTS | PR00412 | Epoxide hydrolase signature | IPR000639 | Epoxide hydrolase-like | 308 | 330 | 1.8E-6 |
| PRINTS | PR00111 | Alpha/beta hydrolase fold signature | IPR000073 | Alpha/beta hydrolase fold-1 | 138 | 151 | 1.5E-11 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.