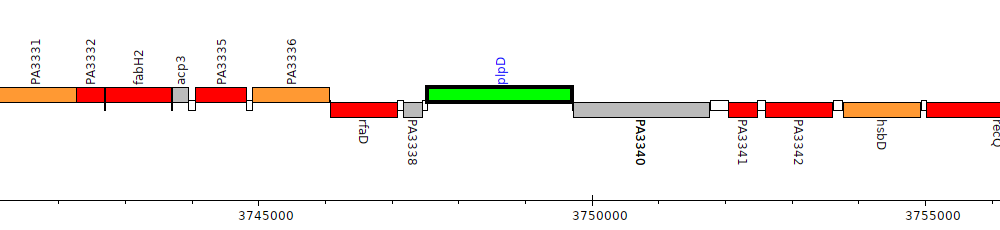
Pseudomonas aeruginosa PAO1, PA3339 (plpD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016298 | lipase activity | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Cellular Component | GO:0009279 | cell outer membrane | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
20192961 | Reviewed by curator |
| Biological Process | GO:0046819 | protein secretion by the type V secretion system | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
20192961 | Reviewed by curator |
| Biological Process | GO:0046819 | protein secretion by the type V secretion system | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Cellular Component | GO:0005576 | extracellular region | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
20192961 | Reviewed by curator |
| Cellular Component | GO:0005576 | extracellular region | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Molecular Function | GO:0016298 | lipase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
20192961 | Reviewed by curator |
| Biological Process | GO:0006629 | lipid metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF01734
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0019867 | outer membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF07244
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Type V secretion system |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Type Vd Secretion System |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.10.20.310 | membrane protein fhac | - | - | 329 | 402 | 1.8E-9 |
| Pfam | PF07244 | Surface antigen variable number repeat | IPR010827 | POTRA domain, BamA/TamA-like | 333 | 393 | 7.0E-5 |
| CDD | cd07205 | Pat_PNPLA6_PNPLA7_NTE1_like | - | - | 25 | 222 | 2.81466E-64 |
| Pfam | PF01103 | Omp85 superfamily domain | IPR000184 | Bacterial surface antigen (D15) | 449 | 728 | 1.2E-9 |
| Gene3D | G3DSA:3.40.1090.10 | Cytosolic phospholipase A2 catalytic domain | - | - | 15 | 126 | 4.4E-31 |
| Gene3D | G3DSA:2.40.160.50 | membrane protein fhac: a member of the omp85/tpsb transporter family | - | - | 411 | 728 | 3.2E-19 |
| SUPERFAMILY | SSF52151 | FabD/lysophospholipase-like | IPR016035 | Acyl transferase/acyl hydrolase/lysophospholipase | 17 | 302 | 2.88E-61 |
| PANTHER | PTHR14226 | NEUROPATHY TARGET ESTERASE/SWISS CHEESE D.MELANOGASTER | - | - | 22 | 317 | 3.4E-55 |
| Gene3D | G3DSA:3.40.1090.10 | Cytosolic phospholipase A2 catalytic domain | - | - | 133 | 269 | 1.1E-25 |
| Pfam | PF01734 | Patatin-like phospholipase | IPR002641 | Patatin-like phospholipase domain | 27 | 219 | 8.9E-28 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.