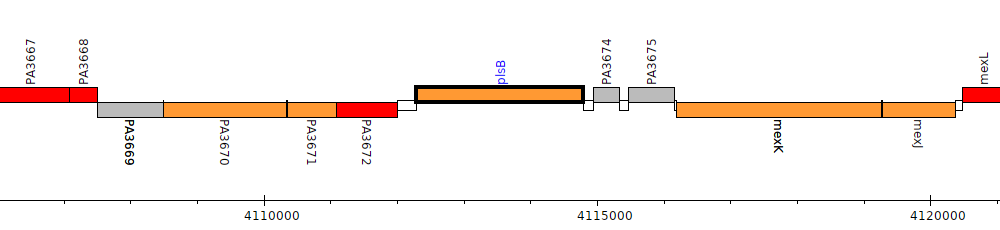
Pseudomonas aeruginosa PAO1, PA3673 (plsB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0044255 | cellular lipid metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0004791 | thioredoxin-disulfide reductase activity | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0044255 | cellular lipid metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR12563
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008374 | O-acyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR12563
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0008654 | phospholipid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03703
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005886 | plasma membrane |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03703
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004366 | glycerol-3-phosphate O-acyltransferase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03703
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016746 | transferase activity, transferring acyl groups |
Inferred from Sequence Model
Term mapped from: InterPro:SM00563
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00564 | Glycerophospholipid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Glycerolipid metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00561 | Glycerolipid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Hamap | MF_00393 | Glycerol-3-phosphate acyltransferase [plsB]. | IPR028354 | Glycerol-3-phosphate acyltransferase, PlsB | 15 | 816 | 31.923655 |
| PIRSF | PIRSF000437 | GPAT_DHAPAT | IPR022284 | Glycerol-3-phosphate O-acyltransferase/Dihydroxyacetone phosphate acyltransferase | 21 | 827 | 0.0 |
| Pfam | PF19277 | Glycerol-3-phosphate acyltransferase C-terminal region | IPR045520 | GPAT/DHAPAT, C-terminal domain | 439 | 782 | 1.6E-25 |
| Pfam | PF01553 | Acyltransferase | IPR002123 | Phospholipid/glycerol acyltransferase | 293 | 428 | 7.4E-20 |
| CDD | cd07993 | LPLAT_DHAPAT-like | IPR041728 | GPAT/DHAPAT, acyltransferase domain | 284 | 485 | 7.45013E-80 |
| NCBIfam | TIGR03703 | JCVI: glycerol-3-phosphate 1-O-acyltransferase PlsB | IPR028354 | Glycerol-3-phosphate acyltransferase, PlsB | 14 | 812 | 0.0 |
| PIRSF | PIRSF500064 | GPAT | IPR028354 | Glycerol-3-phosphate acyltransferase, PlsB | 4 | 829 | 0.0 |
| SUPERFAMILY | SSF69593 | Glycerol-3-phosphate (1)-acyltransferase | - | - | 262 | 523 | 6.93E-32 |
| PANTHER | PTHR12563 | GLYCEROL-3-PHOSPHATE ACYLTRANSFERASE | IPR022284 | Glycerol-3-phosphate O-acyltransferase/Dihydroxyacetone phosphate acyltransferase | 149 | 789 | 0.0 |
| SMART | SM00563 | plsc_2 | IPR002123 | Phospholipid/glycerol acyltransferase | 304 | 431 | 9.4E-27 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.