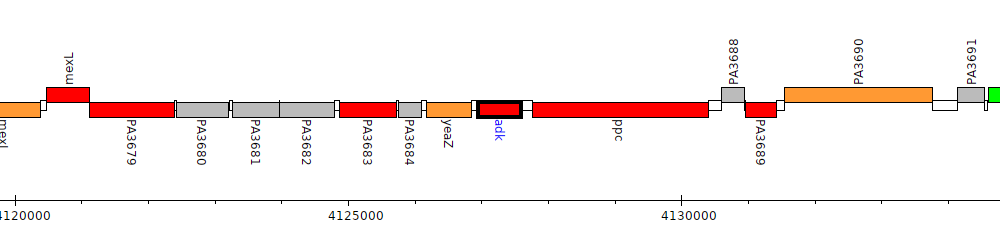
Pseudomonas aeruginosa PAO1, PA3686 (adk)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006163 | purine nucleotide metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0009117 | nucleotide metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0004156 | dihydropteroate synthase activity | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0016776 | phosphotransferase activity, phosphate group as acceptor |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01351
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004017 | adenylate kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01351
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006139 | nucleobase-containing compound metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:cd01428
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd01428
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0019205 | nucleobase-containing compound kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd01428
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae00230 | Purine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00730 | Thiamine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PWY-7219 | adenosine ribonucleotides de novo biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Purine metabolism |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00406 | Adenylate kinase | - | - | 5 | 185 | 1.4E-56 |
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 1 | 212 | 1.01E-43 |
| PRINTS | PR00094 | Adenylate kinase signature | IPR000850 | Adenylate kinase/UMP-CMP kinase | 173 | 187 | 5.1E-36 |
| PANTHER | PTHR23359 | NUCLEOTIDE KINASE | IPR000850 | Adenylate kinase/UMP-CMP kinase | 2 | 210 | 2.0E-59 |
| NCBIfam | TIGR01351 | JCVI: adenylate kinase | IPR006259 | Adenylate kinase subfamily | 2 | 214 | 5.1E-77 |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 1 | 215 | 1.1E-81 |
| PRINTS | PR00094 | Adenylate kinase signature | IPR000850 | Adenylate kinase/UMP-CMP kinase | 32 | 46 | 5.1E-36 |
| PRINTS | PR00094 | Adenylate kinase signature | IPR000850 | Adenylate kinase/UMP-CMP kinase | 4 | 17 | 5.1E-36 |
| Hamap | MF_00235 | Adenylate kinase [adk]. | IPR000850 | Adenylate kinase/UMP-CMP kinase | 1 | 215 | 53.753418 |
| PRINTS | PR00094 | Adenylate kinase signature | IPR000850 | Adenylate kinase/UMP-CMP kinase | 81 | 97 | 5.1E-36 |
| Pfam | PF05191 | Adenylate kinase, active site lid | IPR007862 | Adenylate kinase, active site lid domain | 123 | 158 | 7.3E-18 |
| CDD | cd01428 | ADK | IPR000850 | Adenylate kinase/UMP-CMP kinase | 2 | 206 | 5.08453E-97 |
| PRINTS | PR00094 | Adenylate kinase signature | IPR000850 | Adenylate kinase/UMP-CMP kinase | 156 | 171 | 5.1E-36 |
| FunFam | G3DSA:3.40.50.300:FF:000106 | Adenylate kinase mitochondrial | - | - | 1 | 215 | 4.2E-89 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.