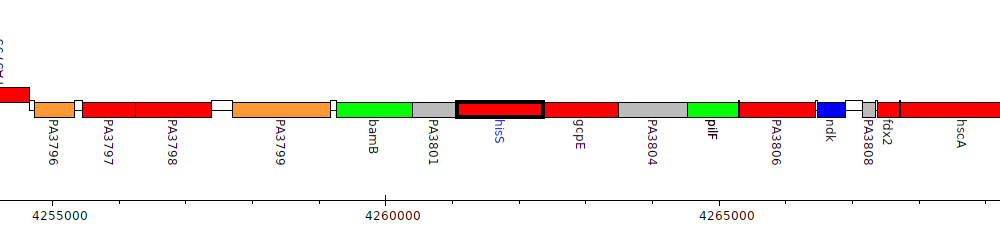
Pseudomonas aeruginosa PAO1, PA3802 (hisS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004814 | arginine-tRNA ligase activity | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0044267 | cellular protein metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00127
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006427 | histidyl-tRNA aminoacylation |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00127
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0005737 | cytoplasm |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR43707
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004821 | histidine-tRNA ligase activity |
Inferred from Sequence Model
Term mapped from: InterPro:MF_00127
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae00970 | Aminoacyl-tRNA biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | TRNA-CHARGING-PWY | tRNA charging | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.800 | - | IPR036621 | Anticodon-binding domain superfamily | 330 | 425 | 6.5E-28 |
| NCBIfam | TIGR00442 | JCVI: histidine--tRNA ligase | IPR015807 | Histidine-tRNA ligase | 6 | 414 | 0.0 |
| CDD | cd00773 | HisRS-like_core | IPR041715 | Class II Histidinyl-tRNA synthetase (HisRS)-like catalytic core domain | 22 | 318 | 4.65759E-97 |
| Pfam | PF13393 | Histidyl-tRNA synthetase | IPR041715 | Class II Histidinyl-tRNA synthetase (HisRS)-like catalytic core domain | 10 | 313 | 4.5E-45 |
| PIRSF | PIRSF001549 | His-tRNA_synth | IPR004516 | Histidine-tRNA ligase/ATP phosphoribosyltransferase regulatory subunit | 1 | 425 | 1.2E-115 |
| CDD | cd00859 | HisRS_anticodon | IPR033656 | Histidyl-anticodon-binding | 330 | 423 | 4.953E-29 |
| PANTHER | PTHR43707 | HISTIDYL-TRNA SYNTHETASE | IPR004516 | Histidine-tRNA ligase/ATP phosphoribosyltransferase regulatory subunit | 4 | 416 | 1.5E-103 |
| SUPERFAMILY | SSF52954 | Class II aaRS ABD-related | - | - | 330 | 425 | 5.4E-23 |
| Pfam | PF03129 | Anticodon binding domain | IPR004154 | Anticodon-binding | 338 | 422 | 2.8E-8 |
| SUPERFAMILY | SSF55681 | Class II aaRS and biotin synthetases | IPR045864 | Class II Aminoacyl-tRNA synthetase/Biotinyl protein ligase (BPL) and lipoyl protein ligase (LPL) | 6 | 323 | 6.7E-102 |
| Gene3D | G3DSA:3.30.930.10 | Bira Bifunctional Protein; Domain 2 | IPR045864 | Class II Aminoacyl-tRNA synthetase/Biotinyl protein ligase (BPL) and lipoyl protein ligase (LPL) | 2 | 326 | 1.2E-116 |
| Hamap | MF_00127 | Histidine--tRNA ligase [hisS]. | IPR015807 | Histidine-tRNA ligase | 3 | 424 | 33.279121 |
| FunFam | G3DSA:3.30.930.10:FF:000005 | Histidine--tRNA ligase | - | - | 2 | 323 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.