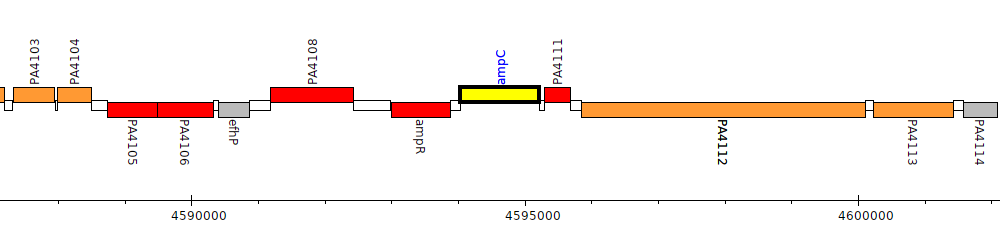
Pseudomonas aeruginosa PAO1, PA4110 (ampC)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0050896 | response to stimulus | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
8405939 | Reviewed by curator |
| Biological Process | GO:0033252 | regulation of beta-lactamase activity | Inferred from Direct Assay | ECO:0000314 direct assay evidence used in manual assertion |
8405939 | Reviewed by curator |
| Molecular Function | GO:0004035 | alkaline phosphatase activity | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01501 | beta-Lactam resistance | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| FunFam | G3DSA:3.40.710.10:FF:000012 | Beta-lactamase | - | - | 29 | 388 | 0.0 |
| SUPERFAMILY | SSF56601 | beta-lactamase/transpeptidase-like | IPR012338 | Beta-lactamase/transpeptidase-like | 32 | 387 | 3.22E-95 |
| PANTHER | PTHR46825 | D-ALANYL-D-ALANINE-CARBOXYPEPTIDASE/ENDOPEPTIDASE AMPH | - | - | 38 | 384 | 6.9E-54 |
| Gene3D | G3DSA:3.40.710.10 | - | IPR012338 | Beta-lactamase/transpeptidase-like | 29 | 388 | 0.0 |
| Pfam | PF00144 | Beta-lactamase | IPR001466 | Beta-lactamase-related | 38 | 385 | 6.7E-80 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.