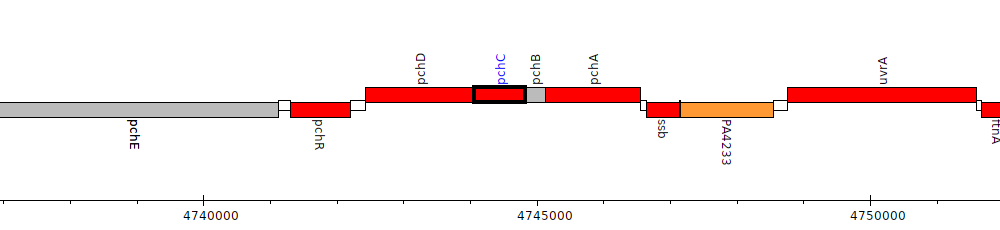
Pseudomonas aeruginosa PAO1, PA4229 (pchC)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0042864 | pyochelin biosynthetic process | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
15375116 | Reviewed by curator |
| Biological Process | GO:0009058 | biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:SM00824
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Pyochelin synthesis |
ECO:0000037
not_recorded |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR11487 | THIOESTERASE | IPR012223 | Thioesterase type II, NRPS/PKS/S-FAS | 4 | 240 | 4.7E-59 |
| Gene3D | G3DSA:3.40.50.1820 | alpha/beta hydrolase | IPR029058 | Alpha/Beta hydrolase fold | 1 | 243 | 6.1E-58 |
| SUPERFAMILY | SSF53474 | alpha/beta-Hydrolases | IPR029058 | Alpha/Beta hydrolase fold | 5 | 240 | 6.1E-46 |
| SMART | SM00824 | Thioesterase | IPR020802 | Polyketide synthase, thioesterase domain | 20 | 236 | 0.0063 |
| Pfam | PF00975 | Thioesterase domain | IPR001031 | Thioesterase | 18 | 237 | 1.6E-60 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.