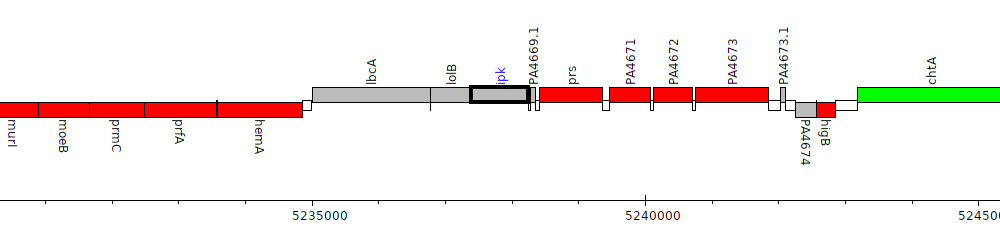
Pseudomonas aeruginosa PAO1, PA4669 (ipk)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0008299 | isoprenoid biosynthetic process | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
21282208 | Reviewed by curator |
| Molecular Function | GO:0050515 | 4-(cytidine 5'-diphospho)-2-C-methyl-D-erythritol kinase activity | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
21282208 | Reviewed by curator |
| Biological Process | GO:0016114 | terpenoid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF010376
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0050515 | 4-(cytidine 5'-diphospho)-2-C-methyl-D-erythritol kinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF010376
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00288
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Terpenoid biosynthesis |
ECO:0000037
not_recorded |
|||
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00900 | Terpenoid backbone biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00288 | GHMP kinases N terminal domain | IPR006204 | GHMP kinase N-terminal domain | 87 | 144 | 1.6E-10 |
| Gene3D | G3DSA:3.30.70.890 | - | IPR036554 | GHMP kinase, C-terminal domain superfamily | 162 | 279 | 2.9E-30 |
| PIRSF | PIRSF010376 | IspE | IPR004424 | 4-diphosphocytidyl-2C-methyl-D-erythritol kinase | 3 | 279 | 4.5E-87 |
| Pfam | PF08544 | GHMP kinases C terminal | IPR013750 | GHMP kinase, C-terminal domain | 205 | 254 | 2.5E-5 |
| SUPERFAMILY | SSF55060 | GHMP Kinase, C-terminal domain | IPR036554 | GHMP kinase, C-terminal domain superfamily | 160 | 277 | 2.73E-33 |
| Gene3D | G3DSA:3.30.230.10 | - | IPR014721 | Small ribosomal subunit protein uS5 domain 2-type fold, subgroup | 1 | 160 | 4.2E-59 |
| SUPERFAMILY | SSF54211 | Ribosomal protein S5 domain 2-like | IPR020568 | Ribosomal protein uS5 domain 2-type superfamily | 7 | 158 | 1.97E-47 |
| NCBIfam | TIGR00154 | JCVI: 4-(cytidine 5'-diphospho)-2-C-methyl-D-erythritol kinase | IPR004424 | 4-diphosphocytidyl-2C-methyl-D-erythritol kinase | 4 | 278 | 2.4E-83 |
| FunFam | G3DSA:3.30.230.10:FF:000022 | 4-diphosphocytidyl-2-C-methyl-D-erythritol kinase | - | - | 1 | 160 | 2.6E-79 |
| Hamap | MF_00061 | Putative 4-diphosphocytidyl-2-C-methyl-D-erythritol kinase [ispE]. | IPR004424 | 4-diphosphocytidyl-2C-methyl-D-erythritol kinase | 5 | 271 | 33.124512 |
| PANTHER | PTHR43527 | 4-DIPHOSPHOCYTIDYL-2-C-METHYL-D-ERYTHRITOL KINASE, CHLOROPLASTIC | - | - | 3 | 262 | 1.8E-71 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.