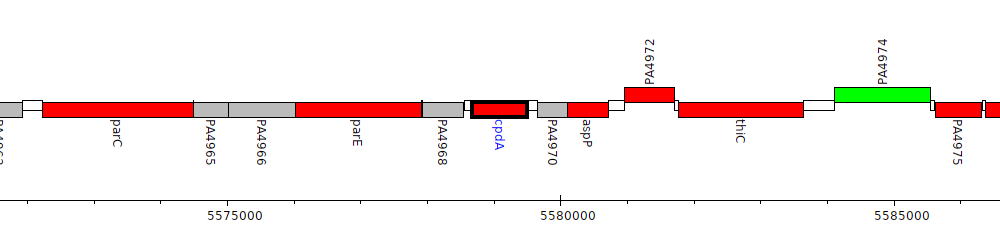
Pseudomonas aeruginosa PAO1, PA4969 (cpdA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Molecular Function | GO:0016787 | hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00149
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004112 | cyclic-nucleotide phosphodiesterase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd07402
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae00230 | Purine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae02025 | Biofilm formation - Pseudomonas aeruginosa | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR42988 | PHOSPHOHYDROLASE | - | - | 14 | 269 | 1.5E-50 |
| CDD | cd07402 | MPP_GpdQ | IPR026575 | Cyclic nucleotide phosphodiesterase GpdQ/CpdA-like | 16 | 250 | 1.21536E-102 |
| Pfam | PF00149 | Calcineurin-like phosphoesterase | IPR004843 | Calcineurin-like phosphoesterase domain, ApaH type | 17 | 203 | 5.5E-13 |
| Gene3D | G3DSA:3.60.21.10 | - | IPR029052 | Metallo-dependent phosphatase-like | 1 | 266 | 1.9E-74 |
| SUPERFAMILY | SSF56300 | Metallo-dependent phosphatases | IPR029052 | Metallo-dependent phosphatase-like | 16 | 267 | 9.3E-56 |
| FunFam | G3DSA:3.60.21.10:FF:000208 | 3',5'-cyclic adenosine monophosphate phosphodiesterase CpdA | - | - | 1 | 266 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.