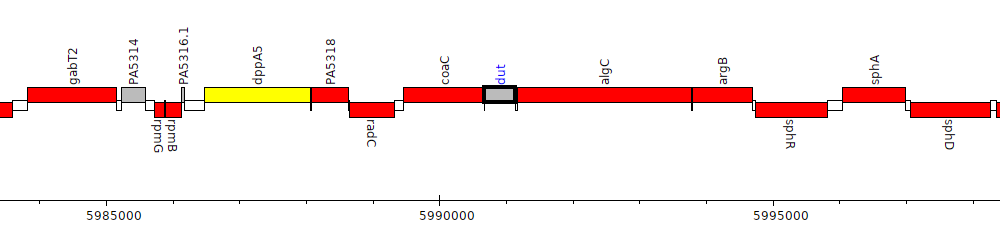
Pseudomonas aeruginosa PAO1, PA5321 (dut)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0005829 | cytosol |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:P06968
|
ECO:0000250 sequence similarity evidence used in manual assertion |
16858726 | Reviewed by curator |
| Molecular Function | GO:0004170 | dUTP diphosphatase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:P06968
|
ECO:0000250 sequence similarity evidence used in manual assertion |
346589 | Reviewed by curator |
| Biological Process | GO:0046081 | dUTP catabolic process |
Inferred from Sequence or Structural Similarity
Term mapped from: UniProtKB:P06968
|
ECO:0000250 sequence similarity evidence used in manual assertion |
346589 | Reviewed by curator |
| Biological Process | GO:0006226 | dUMP biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00576
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0046081 | dUTP catabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00576
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000287 | magnesium ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00576
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004170 | dUTP diphosphatase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00576
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCyc | PWY-7187 | pyrimidine deoxyribonucleotides de novo biosynthesis II | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Pyrimidine metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PWY0-166 | superpathway of pyrimidine deoxyribonucleotides de novo biosynthesis (E. coli) | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00240 | Pyrimidine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PWY-7184 | pyrimidine deoxyribonucleotides de novo biosynthesis I | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd07557 | trimeric_dUTPase | IPR033704 | dUTPase, trimeric | 28 | 121 | 1.20454E-29 |
| NCBIfam | TIGR00576 | JCVI: dUTP diphosphatase | IPR008181 | Deoxyuridine triphosphate nucleotidohydrolase | 10 | 150 | 5.9E-47 |
| FunFam | G3DSA:2.70.40.10:FF:000002 | dUTP diphosphatase | - | - | 1 | 151 | 7.5E-76 |
| Gene3D | G3DSA:2.70.40.10 | - | IPR036157 | dUTPase-like superfamily | 1 | 150 | 2.6E-54 |
| SUPERFAMILY | SSF51283 | dUTPase-like | IPR036157 | dUTPase-like superfamily | 2 | 149 | 1.57E-52 |
| Pfam | PF00692 | dUTPase | IPR029054 | dUTPase-like | 17 | 149 | 5.6E-40 |
| PANTHER | PTHR11241 | DEOXYURIDINE 5'-TRIPHOSPHATE NUCLEOTIDOHYDROLASE | IPR008181 | Deoxyuridine triphosphate nucleotidohydrolase | 11 | 149 | 4.0E-34 |
| Hamap | MF_00116 | Deoxyuridine 5'-triphosphate nucleotidohydrolase [dut]. | IPR008181 | Deoxyuridine triphosphate nucleotidohydrolase | 6 | 150 | 33.191273 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.