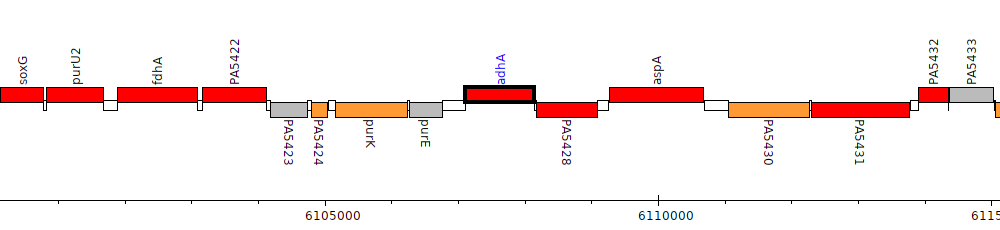
Pseudomonas aeruginosa PAO1, PA5427 (adhA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006631 | fatty acid metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0015976 | carbon utilization | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0004345 | glucose-6-phosphate dehydrogenase activity | Inferred from Direct Assay | ECO:0000002 direct assay evidence |
||
| Biological Process | GO:0044248 | cellular catabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006570 | tyrosine metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006091 | generation of precursor metabolites and energy | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006094 | gluconeogenesis | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0046486 | glycerolipid metabolic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Biological Process | GO:0006096 | glycolytic process | Inferred from Sequence or Structural Similarity | ECO:0000044 sequence similarity evidence |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00829
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| PseudoCAP | Fatty acid metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae00350 | Tyrosine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | FERMENTATION-PWY | mixed acid fermentation | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Bile acid biosynthesis |
ECO:0000037
not_recorded |
|||
| KEGG | pae01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00010 | Glycolysis / Gluconeogenesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Tyrosine metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae00071 | Fatty acid degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00626 | Naphthalene degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | PWY-5480 | pyruvate fermentation to ethanol I | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| KEGG | pae00625 | Chloroalkane and chloroalkene degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCyc | P461-PWY | hexitol fermentation to lactate, formate, ethanol and acetate | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCyc | P122-PWY | heterolactic fermentation | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
| PseudoCAP | Glycolysis / Gluconeogenesis |
ECO:0000037
not_recorded |
|||
| PseudoCAP | Glycerolipid metabolism |
ECO:0000037
not_recorded |
|||
| KEGG | pae01220 | Degradation of aromatic compounds | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00107 | Zinc-binding dehydrogenase | IPR013149 | Alcohol dehydrogenase-like, C-terminal | 181 | 304 | 2.7E-27 |
| CDD | cd08297 | CAD3 | - | - | 7 | 342 | 0.0 |
| SUPERFAMILY | SSF50129 | GroES-like | IPR011032 | GroES-like superfamily | 2 | 340 | 2.74E-58 |
| SMART | SM00829 | PKS_ER_names_mod | IPR020843 | Polyketide synthase, enoylreductase domain | 16 | 340 | 7.7E-6 |
| Gene3D | G3DSA:3.90.180.10 | - | - | - | 12 | 334 | 0.0 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 154 | 291 | 0.0 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 145 | 310 | 1.07E-45 |
| FunFam | G3DSA:3.40.50.720:FF:000039 | Alcohol dehydrogenase AdhP | - | - | 154 | 291 | 3.0E-50 |
| FunFam | G3DSA:3.90.180.10:FF:000002 | Alcohol dehydrogenase AdhP | - | - | 19 | 161 | 8.2E-61 |
| PANTHER | PTHR42940 | ALCOHOL DEHYDROGENASE 1-RELATED | - | - | 1 | 342 | 9.0E-127 |
| Pfam | PF08240 | Alcohol dehydrogenase GroES-like domain | IPR013154 | Alcohol dehydrogenase-like, N-terminal | 31 | 136 | 3.6E-33 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.