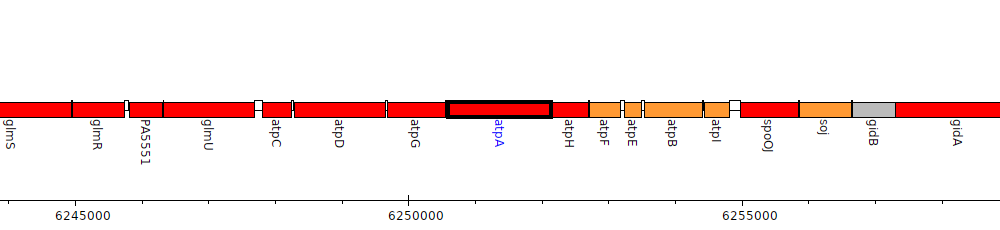
Pseudomonas aeruginosa PAO1, PA5556 (atpA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0045261 | proton-transporting ATP synthase complex, catalytic core F(1) | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
153904 | Reviewed by curator |
| Biological Process | GO:0015986 | ATP synthesis coupled proton transport | Inferred from Sequence or Structural Similarity | ECO:0000250 sequence similarity evidence used in manual assertion |
4256722 | Reviewed by curator |
| Biological Process | GO:0046034 | ATP metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF02874
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01346
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00006
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0045261 | proton-transporting ATP synthase complex, catalytic core F(1) |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01346
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:1902600 | proton transmembrane transport |
Inferred from Sequence Model
Term mapped from: InterPro:PF02874
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0032559 | adenyl ribonucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:cd01132
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0015986 | ATP synthesis coupled proton transport |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01346
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pae01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pae00190 | Oxidative phosphorylation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| PseudoCAP | Oxidative phosphorylation |
ECO:0000037
not_recorded |
|||
| PseudoCyc | PWY-7219 | adenosine ribonucleotides de novo biosynthesis | 19.5 |
ECO:0000250
sequence similarity evidence used in manual assertion |
12867747 |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 97 | 382 | 1.3E-92 |
| Gene3D | G3DSA:2.40.30.20 | - | IPR023366 | ATP synthase subunit alpha, N-terminal domain-like superfamily | 1 | 94 | 1.2E-33 |
| FunFam | G3DSA:3.40.50.300:FF:000002 | ATP synthase subunit alpha | - | - | 95 | 383 | 0.0 |
| Gene3D | G3DSA:1.20.150.20 | - | IPR038376 | ATP synthase, alpha subunit, C-terminal domain superfamily | 384 | 514 | 2.3E-48 |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 95 | 383 | 1.8E-120 |
| Pfam | PF00006 | ATP synthase alpha/beta family, nucleotide-binding domain | IPR000194 | ATPase, F1/V1/A1 complex, alpha/beta subunit, nucleotide-binding domain | 150 | 376 | 2.4E-71 |
| FunFam | G3DSA:2.40.30.20:FF:000001 | ATP synthase subunit alpha | - | - | 1 | 94 | 8.3E-37 |
| SUPERFAMILY | SSF47917 | C-terminal domain of alpha and beta subunits of F1 ATP synthase | - | - | 384 | 512 | 6.02E-45 |
| PANTHER | PTHR48082 | ATP SYNTHASE SUBUNIT ALPHA, MITOCHONDRIAL | IPR005294 | ATP synthase, F1 complex, alpha subunit | 3 | 510 | 0.0 |
| CDD | cd18116 | ATP-synt_F1_alpha_N | - | - | 28 | 94 | 2.35385E-30 |
| Pfam | PF02874 | ATP synthase alpha/beta family, beta-barrel domain | IPR004100 | ATPase, F1/V1/A1 complex, alpha/beta subunit, N-terminal domain | 30 | 93 | 1.6E-15 |
| Pfam | PF00306 | ATP synthase alpha/beta chain, C terminal domain | IPR000793 | ATP synthase, alpha subunit, C-terminal | 383 | 508 | 2.5E-44 |
| PIRSF | PIRSF039088 | F_ATPase_subunit_alpha | IPR005294 | ATP synthase, F1 complex, alpha subunit | 2 | 514 | 0.0 |
| CDD | cd01132 | F1-ATPase_alpha_CD | IPR033732 | ATP synthase, F1 complex, alpha subunit nucleotide-binding domain | 95 | 379 | 0.0 |
| Hamap | MF_01346 | ATP synthase subunit alpha [atpA]. | IPR005294 | ATP synthase, F1 complex, alpha subunit | 2 | 513 | 50.076256 |
| SUPERFAMILY | SSF50615 | N-terminal domain of alpha and beta subunits of F1 ATP synthase | IPR036121 | ATPase, F1/V1/A1 complex, alpha/beta subunit, N-terminal domain superfamily | 11 | 95 | 2.16E-28 |
| NCBIfam | TIGR00962 | JCVI: F0F1 ATP synthase subunit alpha | IPR005294 | ATP synthase, F1 complex, alpha subunit | 4 | 513 | 0.0 |
| CDD | cd18113 | ATP-synt_F1_alpha_C | IPR000793 | ATP synthase, alpha subunit, C-terminal | 387 | 511 | 5.53446E-62 |
| FunFam | G3DSA:1.20.150.20:FF:000001 | ATP synthase subunit alpha | - | - | 384 | 514 | 3.1E-53 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.