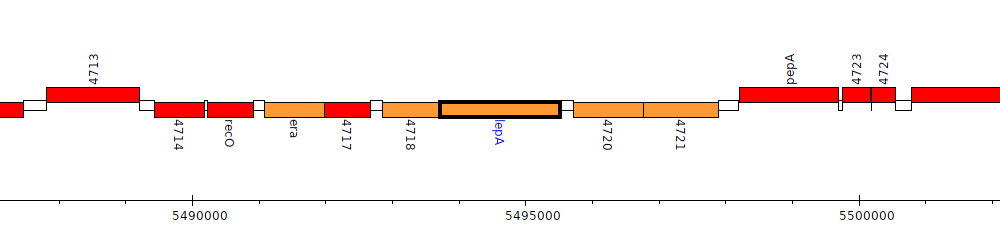
Pseudomonas fluorescens F113, PSF113_4719 (lepA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003924 | GTPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00315
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005525 | GTP binding |
Inferred from Sequence Model
Term mapped from: InterPro:PR00315
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.30.70.2570 | - | IPR038363 | LepA, C-terminal domain superfamily | 482 | 556 | 6.0E-33 |
| PRINTS | PR00315 | GTP-binding elongation factor signature | IPR000795 | Translational (tr)-type GTP-binding domain | 9 | 22 | 1.8E-13 |
| CDD | cd16260 | EF4_III | - | - | 296 | 371 | 5.68362E-47 |
| CDD | cd03699 | EF4_II | - | - | 194 | 279 | 4.53088E-39 |
| Gene3D | G3DSA:2.40.30.10 | Translation factors | - | - | 189 | 287 | 2.6E-32 |
| Pfam | PF00009 | Elongation factor Tu GTP binding domain | IPR000795 | Translational (tr)-type GTP-binding domain | 6 | 184 | 8.9E-51 |
| PRINTS | PR00315 | GTP-binding elongation factor signature | IPR000795 | Translational (tr)-type GTP-binding domain | 93 | 104 | 1.8E-13 |
| Gene3D | G3DSA:3.30.70.240 | - | - | - | 371 | 480 | 4.3E-35 |
| FunFam | G3DSA:3.40.50.300:FF:000078 | Elongation factor 4 | - | - | 4 | 188 | 4.0E-90 |
| CDD | cd03709 | lepA_C | IPR035654 | Elongation factor 4, domain IV | 404 | 482 | 6.71629E-40 |
| CDD | cd01890 | LepA | - | - | 8 | 186 | 7.61617E-126 |
| NCBIfam | TIGR00231 | JCVI: small GTP-binding protein domain | IPR005225 | Small GTP-binding protein domain | 9 | 176 | 4.5E-18 |
| Gene3D | G3DSA:3.40.50.300 | - | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 5 | 188 | 4.2E-61 |
| Pfam | PF06421 | GTP-binding protein LepA C-terminus | IPR013842 | GTP-binding protein LepA, C-terminal | 490 | 596 | 4.3E-50 |
| FunFam | G3DSA:3.30.70.870:FF:000004 | Translation factor GUF1, mitochondrial | - | - | 289 | 370 | 3.2E-37 |
| PANTHER | PTHR43512 | TRANSLATION FACTOR GUF1-RELATED | IPR006297 | Elongation factor 4 | 4 | 596 | 0.0 |
| Pfam | PF00679 | Elongation factor G C-terminus | IPR000640 | Elongation factor EFG, domain V-like | 403 | 487 | 4.1E-19 |
| SUPERFAMILY | SSF54980 | EF-G C-terminal domain-like | IPR035647 | EF-G domain III/V-like | 295 | 373 | 1.94E-18 |
| PRINTS | PR00315 | GTP-binding elongation factor signature | IPR000795 | Translational (tr)-type GTP-binding domain | 77 | 87 | 1.8E-13 |
| SUPERFAMILY | SSF54980 | EF-G C-terminal domain-like | IPR035647 | EF-G domain III/V-like | 403 | 496 | 9.11E-25 |
| Hamap | MF_00071 | Elongation factor 4 [lepA]. | IPR006297 | Elongation factor 4 | 1 | 599 | 301.513947 |
| Pfam | PF03144 | Elongation factor Tu domain 2 | IPR004161 | Translation elongation factor EFTu-like, domain 2 | 208 | 278 | 1.3E-7 |
| FunFam | G3DSA:3.30.70.240:FF:000007 | Translation factor GUF1, mitochondrial | - | - | 371 | 480 | 7.8E-33 |
| FunFam | G3DSA:3.30.70.2570:FF:000001 | Translation factor GUF1, mitochondrial | - | - | 482 | 556 | 3.8E-35 |
| SUPERFAMILY | SSF52540 | P-loop containing nucleoside triphosphate hydrolases | IPR027417 | P-loop containing nucleoside triphosphate hydrolase | 1 | 232 | 2.81E-55 |
| FunFam | G3DSA:2.40.30.10:FF:000015 | Translation factor GUF1, mitochondrial | - | - | 189 | 287 | 1.7E-32 |
| Gene3D | G3DSA:3.30.70.870 | Elongation Factor G (Translational Gtpase), domain 3 | - | - | 289 | 370 | 3.3E-37 |
| PRINTS | PR00315 | GTP-binding elongation factor signature | IPR000795 | Translational (tr)-type GTP-binding domain | 129 | 138 | 1.8E-13 |
| NCBIfam | TIGR01393 | JCVI: translation elongation factor 4 | IPR006297 | Elongation factor 4 | 6 | 597 | 0.0 |
| PRINTS | PR00315 | GTP-binding elongation factor signature | IPR000795 | Translational (tr)-type GTP-binding domain | 52 | 60 | 1.8E-13 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.