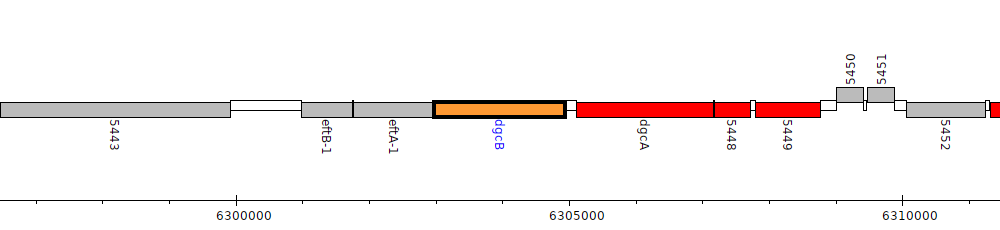
Pseudomonas fluorescens F113, PSF113_5446 (dgcB)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0051536 | iron-sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:1.10.1060.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF13183 | 4Fe-4S dicluster domain | IPR017896 | 4Fe-4S ferredoxin-type, iron-sulphur binding domain | 242 | 332 | 2.0E-11 |
| Gene3D | G3DSA:1.10.1060.10 | - | IPR009051 | Alpha-helical ferredoxin | 219 | 353 | 1.1E-29 |
| SUPERFAMILY | SSF46548 | alpha-helical ferredoxin | - | - | 220 | 348 | 2.27E-21 |
| PANTHER | PTHR43255 | IRON-SULFUR-BINDING OXIDOREDUCTASE FADF-RELATED-RELATED | - | - | 19 | 628 | 0.0 |
| Pfam | PF02754 | Cysteine-rich domain | IPR004017 | Cysteine-rich domain | 521 | 607 | 8.6E-14 |
| Pfam | PF11982 | Domain of unknown function (DUF3483) | IPR021872 | Protein of unknown function DUF3483 | 4 | 220 | 1.4E-88 |
| FunFam | G3DSA:1.10.1060.10:FF:000014 | DgcB, Dimethylglycine catabolism | - | - | 218 | 353 | 7.5E-78 |
| Pfam | PF02754 | Cysteine-rich domain | IPR004017 | Cysteine-rich domain | 398 | 477 | 7.6E-4 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.