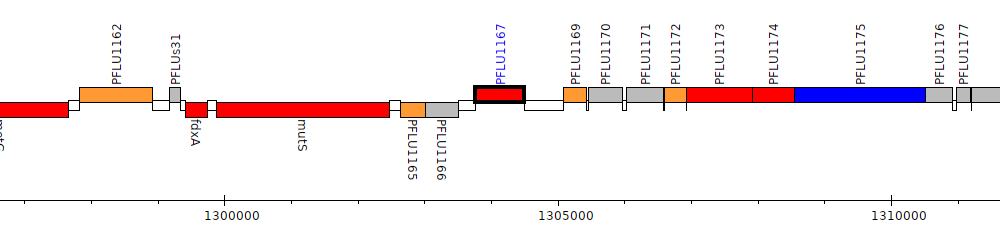
Pseudomonas fluorescens SBW25, PFLU1167
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:1.10.260.40
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00717 | Peptidase S24-like | IPR015927 | Peptidase S24/S26A/S26B/S26C | 112 | 236 | 6.4E-20 |
| Pfam | PF01381 | Helix-turn-helix | IPR001387 | Cro/C1-type helix-turn-helix domain | 10 | 65 | 6.9E-9 |
| CDD | cd00093 | HTH_XRE | IPR001387 | Cro/C1-type helix-turn-helix domain | 10 | 65 | 2.61816E-10 |
| Gene3D | G3DSA:1.10.260.40 | - | IPR010982 | Lambda repressor-like, DNA-binding domain superfamily | 8 | 96 | 3.4E-8 |
| CDD | cd06529 | S24_LexA-like | IPR039418 | LexA-like | 149 | 234 | 2.53E-16 |
| SMART | SM00530 | mbf_short4 | IPR001387 | Cro/C1-type helix-turn-helix domain | 9 | 65 | 1.7E-8 |
| SUPERFAMILY | SSF47413 | lambda repressor-like DNA-binding domains | IPR010982 | Lambda repressor-like, DNA-binding domain superfamily | 6 | 68 | 1.51E-11 |
| Gene3D | G3DSA:2.10.109.10 | Umud Fragment, subunit A | - | - | 104 | 241 | 5.4E-16 |
| SUPERFAMILY | SSF51306 | LexA/Signal peptidase | IPR036286 | LexA/Signal peptidase-like superfamily | 103 | 240 | 3.14E-21 |
| PANTHER | PTHR40661 | - | - | - | 4 | 242 | 2.9E-48 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.