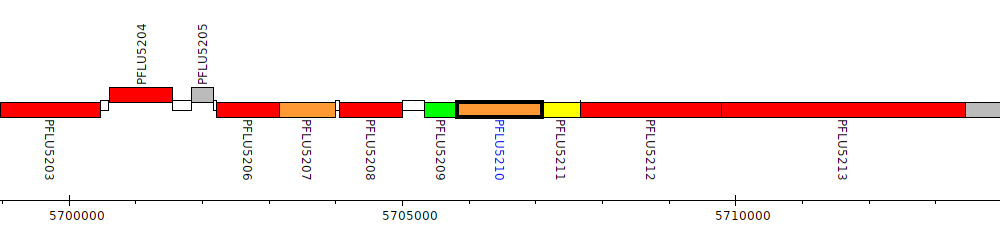
Pseudomonas fluorescens SBW25, PFLU5210
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:SM00304
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0007165 | signal transduction |
Inferred from Sequence Model
Term mapped from: InterPro:SM00304
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM00304 | HAMP_11 | IPR003660 | HAMP domain | 179 | 232 | 1.3E-4 |
| NCBIfam | TIGR00254 | JCVI: diguanylate cyclase (GGDEF) domain | IPR000160 | GGDEF domain | 241 | 399 | 1.6E-43 |
| Pfam | PF17152 | Periplasmic sensor domain | IPR033417 | Periplasmic sensor domain CHASE8 | 41 | 142 | 1.3E-28 |
| SMART | SM00267 | duf1_3 | IPR000160 | GGDEF domain | 234 | 409 | 1.9E-61 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 400 | 420 | - |
| Gene3D | G3DSA:3.30.70.270 | - | IPR043128 | Reverse transcriptase/Diguanylate cyclase domain | 231 | 406 | 5.0E-56 |
| CDD | cd01949 | GGDEF | IPR000160 | GGDEF domain | 246 | 398 | 6.91691E-62 |
| SUPERFAMILY | SSF55073 | Nucleotide cyclase | IPR029787 | Nucleotide cyclase | 250 | 402 | 5.1E-46 |
| PANTHER | PTHR46663 | DIGUANYLATE CYCLASE DGCT-RELATED | - | - | 54 | 409 | 1.2E-72 |
| FunFam | G3DSA:3.30.70.270:FF:000001 | Diguanylate cyclase domain protein | - | - | 240 | 406 | 6.0E-42 |
| Pfam | PF00990 | Diguanylate cyclase, GGDEF domain | IPR000160 | GGDEF domain | 245 | 398 | 1.0E-46 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.