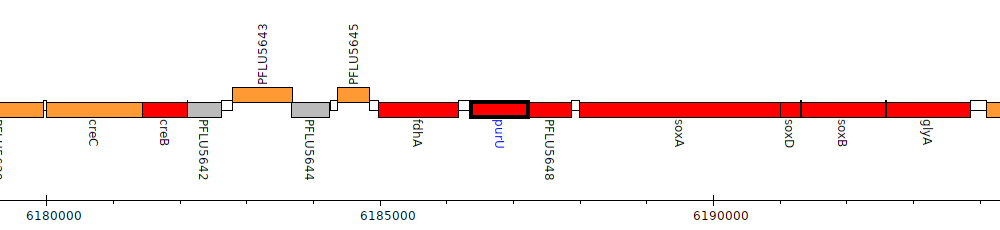
Pseudomonas fluorescens SBW25, PFLU5647 (purU)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0008864 | formyltetrahydrofolate deformylase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR42706
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009058 | biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF00551
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006189 | 'de novo' IMP biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR42706
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| UniPathway | UPA00074 | IMP biosynthesis via de novo pathway | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00630 | Glyoxylate and dicarboxylate metabolism | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00670 | One carbon pool by folate | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
| KEGG (InterPro) | 00970 | Aminoacyl-tRNA biosynthesis | InterPro 5.8-49.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
0 |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR01575 | Formyltetrahydrofolate deformylase signature | IPR004810 | Formyltetrahydrofolate deformylase | 42 | 54 | 8.1E-48 |
| Hamap | MF_01927 | Formyltetrahydrofolate deformylase [purU]. | IPR004810 | Formyltetrahydrofolate deformylase | 5 | 285 | 43.753834 |
| PRINTS | PR01575 | Formyltetrahydrofolate deformylase signature | IPR004810 | Formyltetrahydrofolate deformylase | 91 | 113 | 8.1E-48 |
| PRINTS | PR01575 | Formyltetrahydrofolate deformylase signature | IPR004810 | Formyltetrahydrofolate deformylase | 237 | 259 | 8.1E-48 |
| PRINTS | PR01575 | Formyltetrahydrofolate deformylase signature | IPR004810 | Formyltetrahydrofolate deformylase | 113 | 140 | 8.1E-48 |
| SUPERFAMILY | SSF53328 | Formyltransferase | IPR036477 | Formyl transferase, N-terminal domain superfamily | 91 | 284 | 3.01E-58 |
| NCBIfam | TIGR00655 | JCVI: formyltetrahydrofolate deformylase | IPR004810 | Formyltetrahydrofolate deformylase | 9 | 285 | 2.5E-87 |
| Gene3D | G3DSA:3.40.50.170 | - | - | - | 90 | 285 | 6.3E-56 |
| PRINTS | PR01575 | Formyltetrahydrofolate deformylase signature | IPR004810 | Formyltetrahydrofolate deformylase | 259 | 284 | 8.1E-48 |
| Gene3D | G3DSA:3.30.70.260 | - | - | - | 1 | 88 | 6.8E-24 |
| PANTHER | PTHR42706 | FORMYLTETRAHYDROFOLATE DEFORMYLASE | IPR004810 | Formyltetrahydrofolate deformylase | 6 | 285 | 1.8E-83 |
| Pfam | PF00551 | Formyl transferase | IPR002376 | Formyl transferase, N-terminal | 91 | 266 | 6.3E-37 |
| SUPERFAMILY | SSF55021 | ACT-like | IPR045865 | ACT-like domain | 7 | 84 | 1.37E-8 |
| CDD | cd08648 | FMT_core_Formyl-FH4-Hydrolase_C | IPR041729 | Formyltetrahydrofolate deformylase, C-terminal hydrolase domain | 90 | 285 | 6.77252E-120 |
| PRINTS | PR01575 | Formyltetrahydrofolate deformylase signature | IPR004810 | Formyltetrahydrofolate deformylase | 10 | 36 | 8.1E-48 |
| CDD | cd04875 | ACT_F4HF-DF | IPR044074 | Formyltetrahydrofolate deformylase, ACT domain | 9 | 79 | 1.72613E-24 |
| PIRSF | PIRSF036480 | FormyFH4_hydr | - | - | 5 | 285 | 9.8E-99 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.