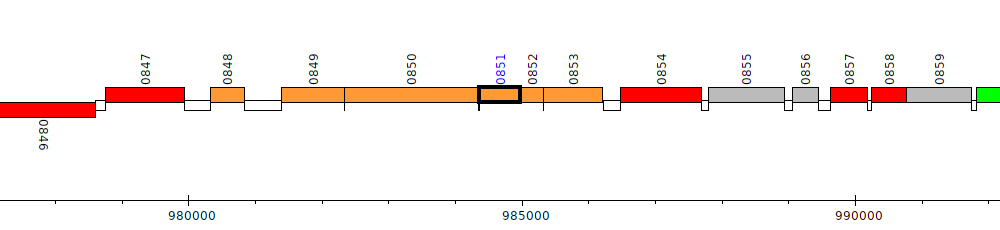
Pseudomonas putida GB-1, PputGB1_0851
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0022904 | respiratory electron transport chain |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:1.20.120.80
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009486 | cytochrome bo3 ubiquinol oxidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02842
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR11403
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0019646 | aerobic electron transport chain |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR11403
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0009055 | electron transfer activity |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR11403
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004129 | cytochrome-c oxidase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00510
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | ppg01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | ppg00190 | Oxidative phosphorylation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd02863 | Ubiquinol_oxidase_III | IPR033946 | Ubiquinol oxidase subunit III domain | 25 | 206 | 9.15749E-92 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 1 | 21 | - |
| FunFam | G3DSA:1.20.120.80:FF:000001 | Cytochrome (Ubi)quinol oxidase subunit III | - | - | 23 | 208 | 4.0E-69 |
| Pfam | PF00510 | Cytochrome c oxidase subunit III | IPR000298 | Cytochrome c oxidase subunit III-like | 27 | 206 | 2.1E-15 |
| Gene3D | G3DSA:1.20.120.80 | - | IPR013833 | Cytochrome c oxidase, subunit III, 4-helical bundle | 23 | 208 | 2.1E-53 |
| NCBIfam | TIGR02842 | JCVI: cytochrome o ubiquinol oxidase subunit III | IPR014206 | Cytochrome o ubiquinol oxidase, subunit III | 29 | 208 | 4.9E-85 |
| PANTHER | PTHR11403 | CYTOCHROME C OXIDASE SUBUNIT III | IPR024791 | Cytochrome c oxidase subunit III | 16 | 206 | 3.6E-40 |
| SUPERFAMILY | SSF81452 | Cytochrome c oxidase subunit III-like | IPR035973 | Cytochrome c oxidase subunit III-like superfamily | 15 | 206 | 7.46E-57 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.