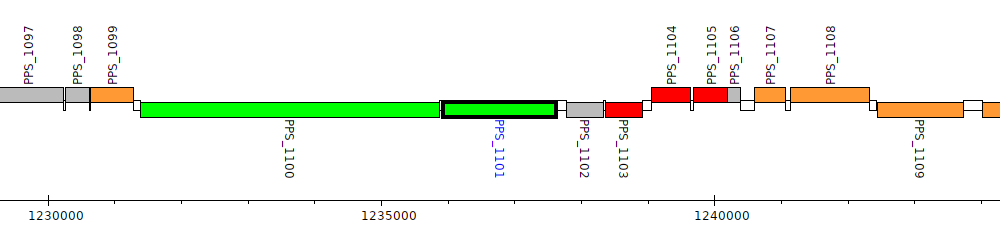
Pseudomonas putida S16, PPS_1101
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0015031 | protein transport |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF029745
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | ppt03070 | Bacterial secretion system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF03865 | Haemolysin secretion/activation protein ShlB/FhaC/HecB | IPR005565 | Haemolysin activator HlyB, C-terminal | 212 | 521 | 4.5E-78 |
| Pfam | PF17287 | POTRA domain | IPR035251 | ShlB, POTRA domain | 155 | 207 | 2.8E-10 |
| PIRSF | PIRSF029745 | FhaC | IPR027282 | Two partner secretion pathway transporter | 5 | 552 | 2.0E-115 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 50 | 72 | - |
| Gene3D | G3DSA:3.10.20.310 | membrane protein fhac | - | - | 64 | 153 | 7.9E-9 |
| Gene3D | G3DSA:2.40.160.50 | membrane protein fhac: a member of the omp85/tpsb transporter family | - | - | 187 | 546 | 1.5E-67 |
| Pfam | PF08479 | POTRA domain, ShlB-type | IPR013686 | Polypeptide-transport-associated, ShlB-type | 78 | 153 | 8.5E-18 |
| PANTHER | PTHR34597 | SLR1661 PROTEIN | - | - | 10 | 531 | 3.0E-85 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.