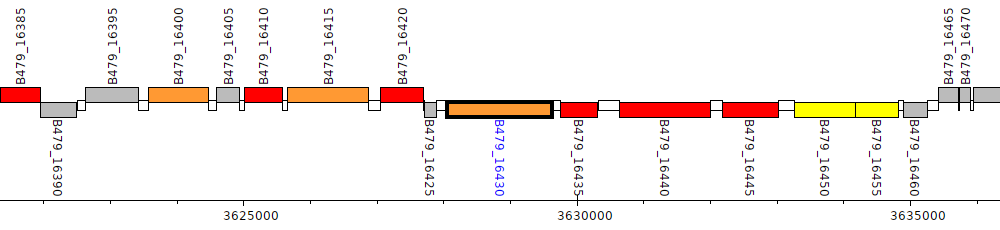
Pseudomonas putida HB3267, B479_16430
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF07992
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0051287 | NAD binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03140
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0050660 | flavin adenine dinucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03140
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0000302 | response to reactive oxygen species |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03140
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0008785 | alkyl hydroperoxide reductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR03140
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 351 | 375 | 1.3E-66 |
| SUPERFAMILY | SSF51905 | FAD/NAD(P)-binding domain | IPR036188 | FAD/NAD(P)-binding domain superfamily | 204 | 512 | 9.88E-53 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 403 | 419 | 1.3E-66 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 213 | 235 | 1.3E-66 |
| PANTHER | PTHR48105 | THIOREDOXIN REDUCTASE 1-RELATED-RELATED | - | - | 211 | 513 | 5.0E-69 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 325 | 448 | 1.2E-97 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 214 | 233 | 1.0E-41 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 245 | 260 | 1.3E-66 |
| Pfam | PF13192 | Thioredoxin domain | IPR012336 | Thioredoxin-like fold | 124 | 194 | 7.4E-8 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 466 | 488 | 1.0E-41 |
| SUPERFAMILY | SSF52833 | Thioredoxin-like | IPR036249 | Thioredoxin-like superfamily | 104 | 195 | 1.12E-21 |
| PIRSF | PIRSF000238 | AhpF | IPR012081 | Alkyl hydroperoxide reductase subunit F | 1 | 519 | 0.0 |
| CDD | cd02974 | AhpF_NTD_N | IPR044142 | AhpF, N-terminal domain, N-terminal TRX-fold subdomain | 1 | 94 | 2.1475E-46 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 313 | 331 | 1.0E-41 |
| Pfam | PF07992 | Pyridine nucleotide-disulphide oxidoreductase | IPR023753 | FAD/NAD(P)-binding domain | 212 | 502 | 1.1E-46 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 265 | 275 | 1.3E-66 |
| SUPERFAMILY | SSF52833 | Thioredoxin-like | IPR036249 | Thioredoxin-like superfamily | 1 | 100 | 8.39E-29 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 440 | 456 | 1.0E-41 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 214 | 509 | 1.2E-97 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 314 | 322 | 1.3E-66 |
| NCBIfam | TIGR03140 | JCVI: alkyl hydroperoxide reductase subunit F | IPR012081 | Alkyl hydroperoxide reductase subunit F | 1 | 515 | 0.0 |
| FunFam | G3DSA:3.50.50.60:FF:000007 | Alkyl hydroperoxide reductase, F subunit | - | - | 325 | 448 | 8.8E-56 |
| CDD | cd03026 | AhpF_NTD_C | IPR044141 | AhpF, N-terminal domain, C-terminal TRX-fold subdomain | 105 | 193 | 4.81841E-53 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 442 | 463 | 1.3E-66 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 476 | 494 | 1.3E-66 |
| Gene3D | G3DSA:3.40.30.80 | - | - | - | 1 | 199 | 2.3E-71 |
| PRINTS | PR00469 | Pyridine nucleotide disulphide reductase class-II signature | - | - | 336 | 348 | 1.3E-66 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 355 | 373 | 1.0E-41 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.