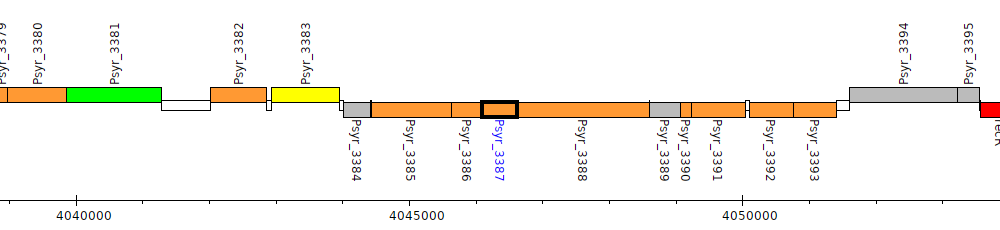
Pseudomonas syringae pv. syringae B728a, Psyr_3387
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0030288 | outer membrane-bounded periplasmic space |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00385
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0015036 | disulfide oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00385
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0017004 | cytochrome complex assembly |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00385
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF08534
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR42852 | THIOL:DISULFIDE INTERCHANGE PROTEIN DSBE | - | - | 39 | 167 | 1.3E-23 |
| Gene3D | G3DSA:3.40.30.10 | Glutaredoxin | - | - | 24 | 177 | 8.6E-45 |
| NCBIfam | TIGR00385 | JCVI: DsbE family thiol:disulfide interchange protein | IPR004799 | Periplasmic protein thiol:disulphide oxidoreductase DsbE | 6 | 169 | 2.3E-54 |
| Pfam | PF08534 | Redoxin | IPR013740 | Redoxin | 37 | 157 | 4.1E-16 |
| CDD | cd03010 | TlpA_like_DsbE | IPR004799 | Periplasmic protein thiol:disulphide oxidoreductase DsbE | 38 | 162 | 1.01256E-69 |
| SUPERFAMILY | SSF52833 | Thioredoxin-like | IPR036249 | Thioredoxin-like superfamily | 41 | 171 | 3.11E-35 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.