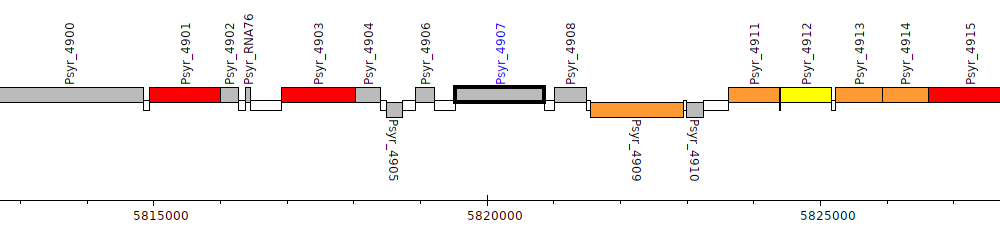
Pseudomonas syringae pv. syringae B728a, Psyr_4907
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF00015
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0007165 | signal transduction |
Inferred from Sequence Model
Term mapped from: InterPro:PF00015
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006935 | chemotaxis |
Inferred from Sequence Model
Term mapped from: InterPro:PR00260
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004888 | transmembrane signaling receptor activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00260
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005515 | protein binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF08447
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | psb02030 | Bacterial chemotaxis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | psb02020 | Two-component system | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00015 | Methyl-accepting chemotaxis protein (MCP) signalling domain | IPR004089 | Methyl-accepting chemotaxis protein (MCP) signalling domain | 275 | 438 | 6.8E-38 |
| PRINTS | PR00260 | Bacterial chemotaxis sensory transducer signature | IPR004090 | Chemotaxis methyl-accepting receptor | 326 | 353 | 2.5E-20 |
| Pfam | PF08447 | PAS fold | IPR013655 | PAS fold-3 | 43 | 128 | 8.6E-18 |
| PANTHER | PTHR32089 | METHYL-ACCEPTING CHEMOTAXIS PROTEIN MCPB | - | - | 251 | 441 | 2.8E-51 |
| NCBIfam | TIGR00229 | JCVI: PAS domain S-box protein | IPR000014 | PAS domain | 153 | 263 | 5.8E-14 |
| Gene3D | G3DSA:3.30.450.20 | PAS domain | - | - | 6 | 126 | 3.2E-19 |
| SUPERFAMILY | SSF55785 | PYP-like sensor domain (PAS domain) | IPR035965 | PAS domain superfamily | 30 | 132 | 2.42E-24 |
| SMART | SM00283 | MA_2 | IPR004089 | Methyl-accepting chemotaxis protein (MCP) signalling domain | 246 | 447 | 2.8E-23 |
| SUPERFAMILY | SSF58104 | Methyl-accepting chemotaxis protein (MCP) signaling domain | - | - | 249 | 441 | 2.88E-55 |
| CDD | cd00130 | PAS | IPR000014 | PAS domain | 152 | 254 | 4.7167E-8 |
| SUPERFAMILY | SSF55785 | PYP-like sensor domain (PAS domain) | IPR035965 | PAS domain superfamily | 153 | 254 | 4.19E-22 |
| Gene3D | G3DSA:1.10.287.950 | - | - | - | 248 | 444 | 8.5E-46 |
| CDD | cd11386 | MCP_signal | - | - | 275 | 438 | 5.5303E-49 |
| PRINTS | PR00260 | Bacterial chemotaxis sensory transducer signature | IPR004090 | Chemotaxis methyl-accepting receptor | 355 | 384 | 2.5E-20 |
| Pfam | PF08447 | PAS fold | IPR013655 | PAS fold-3 | 165 | 250 | 3.3E-15 |
| Gene3D | G3DSA:3.30.450.20 | PAS domain | - | - | 127 | 247 | 3.9E-19 |
| CDD | cd00130 | PAS | IPR000014 | PAS domain | 30 | 132 | 5.84713E-12 |
| SMART | SM00086 | pac_2 | IPR001610 | PAC motif | 215 | 257 | 0.039 |
| PRINTS | PR00260 | Bacterial chemotaxis sensory transducer signature | IPR004090 | Chemotaxis methyl-accepting receptor | 253 | 282 | 2.5E-20 |
| SMART | SM00086 | pac_2 | IPR001610 | PAC motif | 93 | 135 | 0.0031 |
| NCBIfam | TIGR00229 | JCVI: PAS domain S-box protein | IPR000014 | PAS domain | 23 | 141 | 2.8E-15 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.