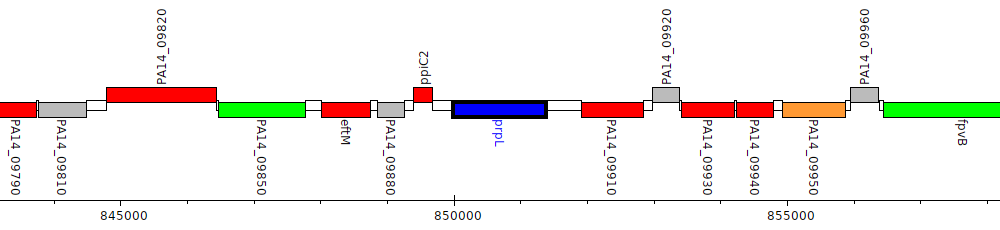
Pseudomonas aeruginosa UCBPP-PA14, PA14_09900 (prpL)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006554 | lysine catabolic process |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA4175
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
11237336 | |
| Molecular Function | GO:0008236 | serine-type peptidase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA4175
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
11237336 |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:2.40.10.10 | - | IPR043504 | Peptidase S1, PA clan, chymotrypsin-like fold | 246 | 349 | 7.0E-46 |
| Gene3D | G3DSA:2.40.10.10 | - | IPR043504 | Peptidase S1, PA clan, chymotrypsin-like fold | 221 | 449 | 7.0E-46 |
| PANTHER | PTHR36234 | LYSYL ENDOPEPTIDASE | - | - | 33 | 458 | 2.1E-79 |
| SUPERFAMILY | SSF50494 | Trypsin-like serine proteases | IPR009003 | Peptidase S1, PA clan | 222 | 459 | 2.71E-34 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.