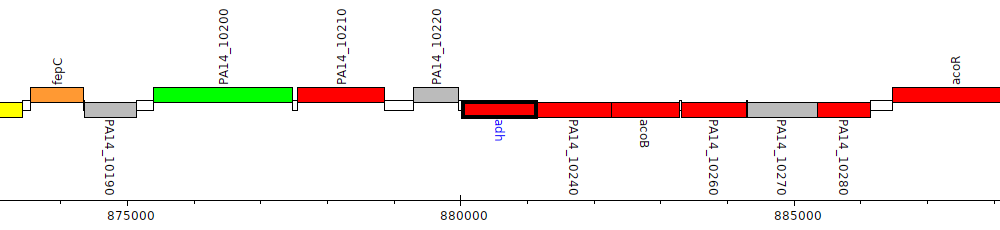
Pseudomonas aeruginosa UCBPP-PA14, PA14_10230 (adh)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:SM00829
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau00650 | Butanoate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 170 | 306 | 4.1E-103 |
| Pfam | PF08240 | Alcohol dehydrogenase GroES-like domain | IPR013154 | Alcohol dehydrogenase-like, N-terminal | 35 | 151 | 1.0E-31 |
| SMART | SM00829 | PKS_ER_names_mod | IPR020843 | Polyketide synthase, enoylreductase domain | 18 | 359 | 1.9E-4 |
| CDD | cd08233 | butanediol_DH_like | - | - | 11 | 360 | 0.0 |
| SUPERFAMILY | SSF50129 | GroES-like | IPR011032 | GroES-like superfamily | 9 | 192 | 9.13E-51 |
| PANTHER | PTHR43161 | SORBITOL DEHYDROGENASE | - | - | 12 | 360 | 8.0E-92 |
| Gene3D | G3DSA:3.90.180.10 | - | - | - | 20 | 341 | 4.1E-103 |
| Pfam | PF13823 | Alcohol dehydrogenase GroES-associated | IPR027399 | Alcohol dehydrogenase GroES-associated | 11 | 28 | 2.4E-6 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 159 | 315 | 8.97E-39 |
| Pfam | PF00107 | Zinc-binding dehydrogenase | IPR013149 | Alcohol dehydrogenase-like, C-terminal | 194 | 320 | 8.1E-28 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.