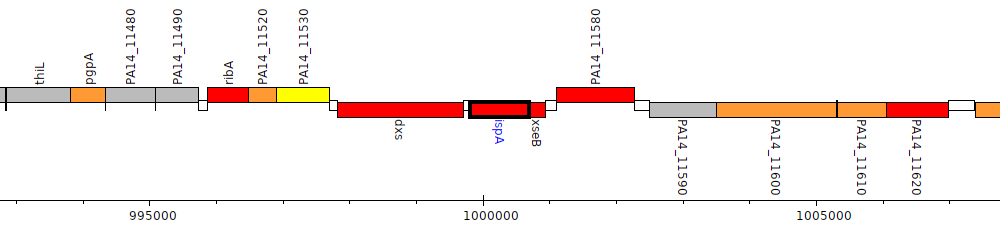
Pseudomonas aeruginosa UCBPP-PA14, PA14_11560 (ispA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0008299 | isoprenoid biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:cd00685
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00900 | Terpenoid backbone biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF48576 | Terpenoid synthases | IPR008949 | Isoprenoid synthase domain superfamily | 5 | 287 | 1.15E-86 |
| SFLD | SFLDS00005 | Isoprenoid Synthase Type I | - | - | 5 | 293 | 0.0 |
| CDD | cd00685 | Trans_IPPS_HT | IPR000092 | Polyprenyl synthetase | 26 | 293 | 2.7311E-73 |
| Pfam | PF00348 | Polyprenyl synthetase | IPR000092 | Polyprenyl synthetase | 29 | 259 | 4.7E-58 |
| SFLD | SFLDG01017 | Polyprenyl Transferase Like | - | - | 5 | 293 | 0.0 |
| Gene3D | G3DSA:1.10.600.10 | Farnesyl Diphosphate Synthase | IPR008949 | Isoprenoid synthase domain superfamily | 1 | 295 | 4.6E-107 |
| PANTHER | PTHR43281 | FARNESYL DIPHOSPHATE SYNTHASE | - | - | 4 | 294 | 7.7E-95 |
| FunFam | G3DSA:1.10.600.10:FF:000001 | Geranylgeranyl diphosphate synthase | - | - | 1 | 295 | 1.4E-115 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.