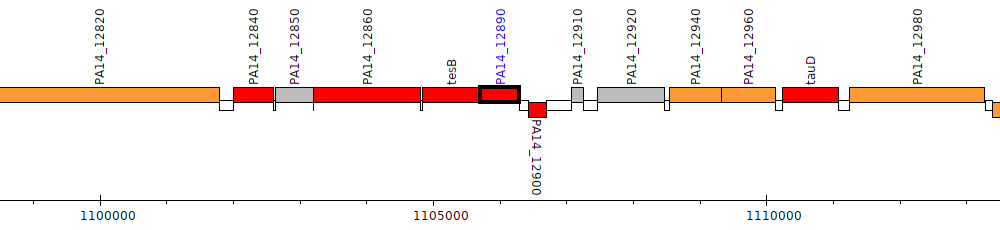
Pseudomonas aeruginosa UCBPP-PA14, PA14_12890
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| CDD | cd01427 | HAD_like | - | - | 75 | 170 | 6.7479E-18 |
| Gene3D | G3DSA:1.10.260.80 | - | - | - | 19 | 69 | 3.3E-80 |
| Gene3D | G3DSA:3.40.50.1000 | - | IPR023214 | HAD superfamily | 8 | 189 | 3.3E-80 |
| Pfam | PF13419 | Haloacid dehalogenase-like hydrolase | IPR041492 | Haloacid dehalogenase-like hydrolase | 41 | 170 | 1.5E-16 |
| NCBIfam | TIGR01509 | JCVI: HAD-IA family hydrolase | IPR006439 | HAD hydrolase, subfamily IA | 8 | 170 | 1.4E-9 |
| SUPERFAMILY | SSF56784 | HAD-like | IPR036412 | HAD-like superfamily | 5 | 195 | 1.81E-37 |
| PANTHER | PTHR43885 | HALOACID DEHALOGENASE-LIKE HYDROLASE | - | - | 5 | 171 | 2.3E-32 |
| NCBIfam | TIGR01549 | JCVI: HAD-IA family hydrolase | IPR006439 | HAD hydrolase, subfamily IA | 10 | 164 | 2.6E-8 |
| SFLD | SFLDG01129 | C1.5: HAD, Beta-PGM, Phosphatase Like | - | - | 6 | 196 | 3.0E-28 |
| SFLD | SFLDS00003 | Haloacid Dehalogenase | - | - | 6 | 196 | 3.0E-28 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.