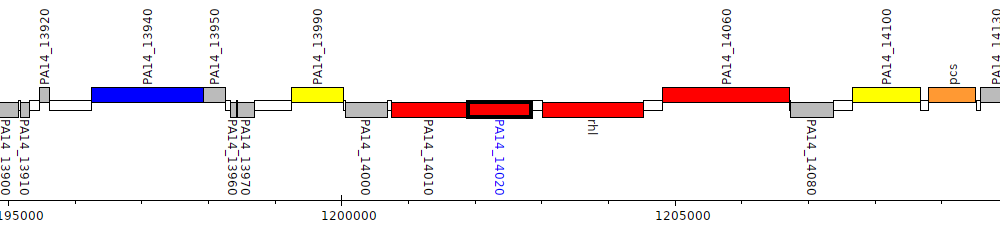
Pseudomonas aeruginosa UCBPP-PA14, PA14_14020
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00472 | D-Arginine and D-ornithine metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 6 | 313 | 4.44E-68 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 126 | 296 | 4.6E-63 |
| PIRSF | PIRSF001439 | CryM | IPR003462 | Ornithine cyclodeaminase/mu-crystallin | 1 | 315 | 3.7E-52 |
| Pfam | PF02423 | Ornithine cyclodeaminase/mu-crystallin family | IPR003462 | Ornithine cyclodeaminase/mu-crystallin | 8 | 309 | 3.9E-33 |
| Gene3D | G3DSA:3.30.1780.10 | ornithine cyclodeaminase, domain 1 | IPR023401 | Ornithine cyclodeaminase, N-terminal | 23 | 310 | 4.6E-63 |
| PANTHER | PTHR13812 | KETIMINE REDUCTASE MU-CRYSTALLIN | IPR003462 | Ornithine cyclodeaminase/mu-crystallin | 7 | 312 | 3.7E-45 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.