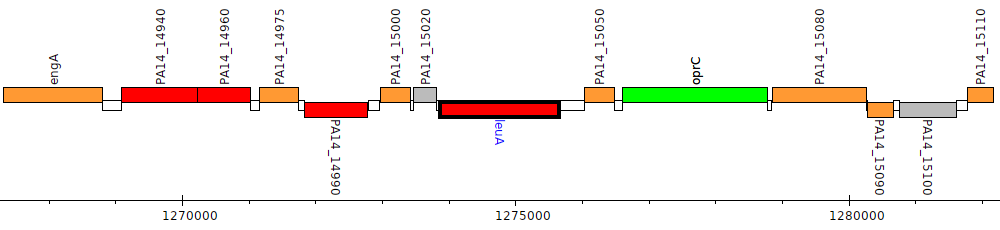
Pseudomonas aeruginosa UCBPP-PA14, PA14_15030 (leuA)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00682
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003852 | 2-isopropylmalate synthase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF08502
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0009098 | leucine biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PF08502
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01230 | Biosynthesis of amino acids | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00620 | Pyruvate metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau00290 | Valine, leucine and isoleucine biosynthesis | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pau01210 | 2-Oxocarboxylic acid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| NCBIfam | TIGR00970 | JCVI: 2-isopropylmalate synthase | IPR005668 | 2-isopropylmalate synthase | 43 | 584 | 0.0 |
| CDD | cd07942 | DRE_TIM_LeuA | IPR039371 | LeuA, N-terminal catalytic TIM barrel domain | 69 | 351 | 0.0 |
| SUPERFAMILY | SSF89000 | post-HMGL domain-like | - | - | 424 | 473 | 1.37E-13 |
| Pfam | PF00682 | HMGL-like | IPR000891 | Pyruvate carboxyltransferase | 69 | 349 | 7.8E-79 |
| SUPERFAMILY | SSF110921 | 2-isopropylmalate synthase LeuA, allosteric (dimerisation) domain | IPR036230 | 2-isopropylmalate synthase LeuA, allosteric (dimerisation) domain superfamily | 479 | 583 | 2.22E-33 |
| Pfam | PF08502 | LeuA allosteric (dimerisation) domain | IPR013709 | 2-isopropylmalate synthase LeuA, allosteric (dimerisation) domain | 458 | 583 | 9.0E-29 |
| SUPERFAMILY | SSF51569 | Aldolase | - | - | 58 | 366 | 1.83E-98 |
| PANTHER | PTHR46911 | - | - | - | 38 | 590 | 0.0 |
| Hamap | MF_00572 | 2-isopropylmalate synthase [leuA]. | IPR005668 | 2-isopropylmalate synthase | 44 | 585 | 54.365555 |
| FunFam | G3DSA:3.20.20.70:FF:000045 | 2-isopropylmalate synthase | - | - | 47 | 412 | 0.0 |
| SMART | SM00917 | LeuA_dimer_2 | IPR013709 | 2-isopropylmalate synthase LeuA, allosteric (dimerisation) domain | 458 | 584 | 7.7E-31 |
| FunFam | G3DSA:3.30.160.270:FF:000006 | 2-isopropylmalate synthase | - | - | 413 | 583 | 2.2E-73 |
| Gene3D | G3DSA:3.20.20.70 | Aldolase class I | IPR013785 | Aldolase-type TIM barrel | 48 | 412 | 0.0 |
| Gene3D | G3DSA:3.30.160.270 | - | IPR036230 | 2-isopropylmalate synthase LeuA, allosteric (dimerisation) domain superfamily | 413 | 583 | 6.4E-56 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.