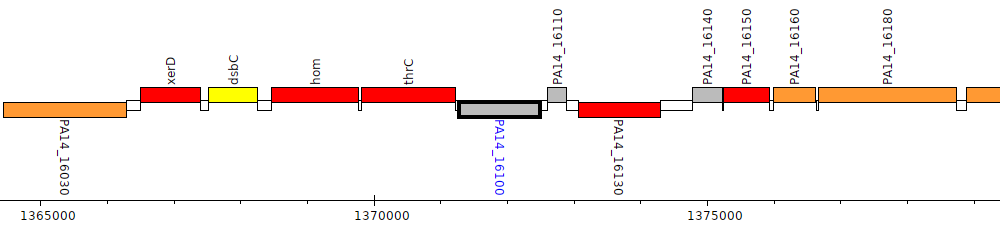
Pseudomonas aeruginosa UCBPP-PA14, PA14_16100
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0050525 | cutinase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF12740
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0052689 | carboxylic ester hydrolase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF12740
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.1820 | alpha/beta hydrolase | IPR029058 | Alpha/Beta hydrolase fold | 149 | 392 | 3.0E-37 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 16 | 37 | - |
| SUPERFAMILY | SSF53474 | alpha/beta-Hydrolases | IPR029058 | Alpha/Beta hydrolase fold | 159 | 371 | 5.26E-19 |
| PANTHER | PTHR33428 | CHLOROPHYLLASE-2, CHLOROPLASTIC | - | - | 142 | 377 | 1.3E-27 |
| Pfam | PF12740 | Chlorophyllase enzyme | IPR041127 | Cutinase | 158 | 288 | 2.1E-8 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.