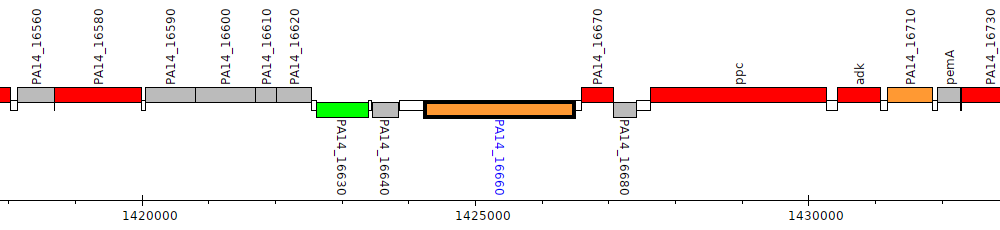
Pseudomonas aeruginosa UCBPP-PA14, PA14_16660
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005215 | transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016887 | ATPase activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0000166 | nucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.40.1110.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PR00941
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0019829 | ATPase-coupled cation transmembrane transporter activity |
Inferred from Sequence Model
Term mapped from: InterPro:PR00941
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0046872 | metal ion binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF55008
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR01494
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006812 | cation transport |
Inferred from Sequence Model
Term mapped from: InterPro:PR00941
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR48085 | CADMIUM/ZINC-TRANSPORTING ATPASE HMA2-RELATED | - | - | 43 | 740 | 0.0 |
| CDD | cd00371 | HMA | IPR006121 | Heavy metal-associated domain, HMA | 47 | 107 | 7.45014E-5 |
| SUPERFAMILY | SSF81665 | Calcium ATPase, transmembrane domain M | IPR023298 | P-type ATPase, transmembrane domain superfamily | 185 | 422 | 3.27E-14 |
| Gene3D | G3DSA:3.30.70.100 | - | - | - | 39 | 115 | 5.8E-5 |
| SFLD | SFLDS00003 | Haloacid Dehalogenase | - | - | 417 | 686 | 4.764E-44 |
| PRINTS | PR00941 | Cadmium-transporting ATPase signature | IPR027256 | P-type ATPase, subfamily IB | 219 | 239 | 1.6E-10 |
| PRINTS | PR00941 | Cadmium-transporting ATPase signature | IPR027256 | P-type ATPase, subfamily IB | 300 | 316 | 1.6E-10 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 1 | 26 | - |
| NCBIfam | TIGR01494 | JCVI: HAD-IC family P-type ATPase | IPR001757 | P-type ATPase | 207 | 460 | 2.3E-36 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 284 | 298 | 6.6E-21 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 435 | 449 | 6.6E-21 |
| Gene3D | G3DSA:2.70.150.10 | - | - | - | 228 | 339 | 1.5E-30 |
| CDD | cd07545 | P-type_ATPase_Cd-like | - | - | 143 | 739 | 0.0 |
| NCBIfam | TIGR01525 | JCVI: heavy metal translocating P-type ATPase | IPR027256 | P-type ATPase, subfamily IB | 184 | 737 | 0.0 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 634 | 653 | 6.6E-21 |
| SUPERFAMILY | SSF55008 | HMA, heavy metal-associated domain | IPR036163 | Heavy metal-associated domain superfamily | 43 | 113 | 6.17E-10 |
| Pfam | PF00702 | haloacid dehalogenase-like hydrolase | - | - | 433 | 648 | 2.9E-41 |
| NCBIfam | TIGR01494 | JCVI: HAD-IC family P-type ATPase | IPR001757 | P-type ATPase | 534 | 710 | 4.7E-38 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 1 | 15 | - |
| FunFam | G3DSA:2.70.150.10:FF:000002 | Copper-transporting ATPase 1, putative | - | - | 222 | 338 | 1.2E-32 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 581 | 591 | 6.6E-21 |
| Gene3D | G3DSA:3.40.50.1000 | - | IPR023214 | HAD superfamily | 429 | 685 | 2.3E-81 |
| SFLD | SFLDF00027 | p-type atpase | IPR044492 | P-type ATPase, haloacid dehalogenase domain | 417 | 686 | 4.764E-44 |
| Gene3D | G3DSA:3.40.1110.10 | - | IPR023299 | P-type ATPase, cytoplasmic domain N | 446 | 567 | 2.3E-81 |
| PRINTS | PR00941 | Cadmium-transporting ATPase signature | IPR027256 | P-type ATPase, subfamily IB | 319 | 333 | 1.6E-10 |
| Pfam | PF00122 | E1-E2 ATPase | - | - | 235 | 414 | 3.4E-54 |
| PRINTS | PR00119 | P-type cation-transporting ATPase superfamily signature | - | - | 658 | 670 | 6.6E-21 |
| SUPERFAMILY | SSF56784 | HAD-like | IPR036412 | HAD-like superfamily | 434 | 734 | 5.26E-52 |
| SUPERFAMILY | SSF81653 | Calcium ATPase, transduction domain A | IPR008250 | P-type ATPase, A domain superfamily | 237 | 334 | 2.48E-22 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.