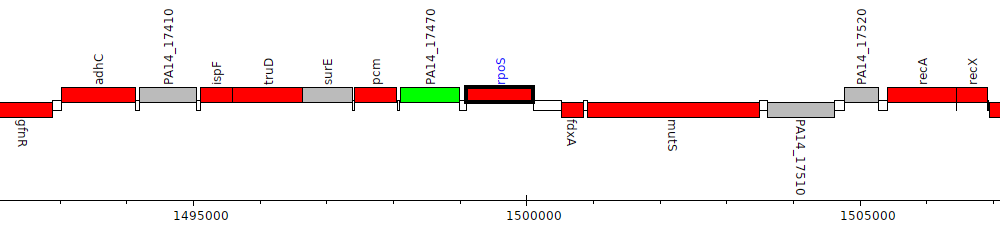
Pseudomonas aeruginosa UCBPP-PA14, PA14_17480 (rpoS)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:2000145 | regulation of cell motility |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA3622
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
10383954 | |
| Biological Process | GO:1900034 | regulation of cellular response to heat |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA3622
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
10383954 | |
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA3622
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
10383954 | |
| Biological Process | GO:1901000 | regulation of response to salt stress |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA3622
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
10383954 | |
| Biological Process | GO:1900407 | regulation of cellular response to oxidative stress |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA3622
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
10383954 | |
| Biological Process | GO:0050921 | positive regulation of chemotaxis |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA3622
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
15687221 | |
| Biological Process | GO:1900233 | positive regulation of single-species biofilm formation on inanimate substrate |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA3622
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
20735777 | |
| Biological Process | GO:1900377 | negative regulation of secondary metabolite biosynthetic process |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA3622
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
10383954 | |
| Molecular Function | GO:0016987 | sigma factor activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00140
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006355 | regulation of transcription, DNA-templated |
Inferred from Sequence Model
Term mapped from: InterPro:PF00140
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003677 | DNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00140
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003700 | DNA-binding transcription factor activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00140
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006352 | DNA-templated transcription, initiation |
Inferred from Sequence Model
Term mapped from: InterPro:PF00140
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PRINTS | PR00046 | Major sigma-70 factor signature | IPR000943 | RNA polymerase sigma-70 | 147 | 155 | 5.9E-28 |
| Pfam | PF04542 | Sigma-70 region 2 | IPR007627 | RNA polymerase sigma-70 region 2 | 99 | 168 | 9.4E-22 |
| NCBIfam | TIGR02394 | JCVI: RNA polymerase sigma factor RpoS | IPR012761 | RNA polymerase sigma factor RpoS | 54 | 333 | 0.0 |
| SUPERFAMILY | SSF88946 | Sigma2 domain of RNA polymerase sigma factors | IPR013325 | RNA polymerase sigma factor, region 2 | 61 | 169 | 2.88E-45 |
| FunFam | G3DSA:1.10.10.10:FF:000046 | RNA polymerase sigma factor RpoS | - | - | 172 | 248 | 7.0E-45 |
| Gene3D | G3DSA:1.10.10.10 | - | IPR036388 | Winged helix-like DNA-binding domain superfamily | 249 | 324 | 2.6E-22 |
| PRINTS | PR00046 | Major sigma-70 factor signature | IPR000943 | RNA polymerase sigma-70 | 307 | 318 | 5.9E-28 |
| Gene3D | G3DSA:1.10.601.10 | RNA Polymerase Primary Sigma Factor | - | - | 26 | 168 | 3.6E-47 |
| Gene3D | G3DSA:1.10.10.10 | - | IPR036388 | Winged helix-like DNA-binding domain superfamily | 172 | 247 | 3.7E-15 |
| Hamap | MF_00959 | RNA polymerase sigma factor RpoS [rpoS]. | IPR012761 | RNA polymerase sigma factor RpoS | 4 | 333 | 40.747299 |
| PRINTS | PR00046 | Major sigma-70 factor signature | IPR000943 | RNA polymerase sigma-70 | 271 | 283 | 5.9E-28 |
| CDD | cd06171 | Sigma70_r4 | - | - | 262 | 319 | 4.7209E-11 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 32 | 50 | - |
| PRINTS | PR00046 | Major sigma-70 factor signature | IPR000943 | RNA polymerase sigma-70 | 123 | 136 | 5.9E-28 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 21 | 50 | - |
| SUPERFAMILY | SSF88659 | Sigma3 and sigma4 domains of RNA polymerase sigma factors | IPR013324 | RNA polymerase sigma factor, region 3/4-like | 230 | 327 | 2.62E-25 |
| FunFam | G3DSA:1.10.601.10:FF:000001 | RNA polymerase sigma factor SigA | - | - | 91 | 171 | 1.1E-46 |
| Pfam | PF04539 | Sigma-70 region 3 | IPR007624 | RNA polymerase sigma-70 region 3 | 179 | 253 | 1.5E-20 |
| NCBIfam | TIGR02937 | JCVI: sigma-70 family RNA polymerase sigma factor | IPR014284 | RNA polymerase sigma-70 like domain | 96 | 322 | 1.1E-36 |
| Pfam | PF04545 | Sigma-70, region 4 | IPR007630 | RNA polymerase sigma-70 region 4 | 267 | 320 | 1.1E-15 |
| PANTHER | PTHR30603 | RNA POLYMERASE SIGMA FACTOR RPO | - | - | 95 | 328 | 6.3E-85 |
| SUPERFAMILY | SSF88659 | Sigma3 and sigma4 domains of RNA polymerase sigma factors | IPR013324 | RNA polymerase sigma factor, region 3/4-like | 172 | 246 | 5.67E-14 |
| FunFam | G3DSA:1.10.10.10:FF:000044 | RNA polymerase sigma factor RpoS | - | - | 250 | 323 | 1.6E-38 |
| PRINTS | PR00046 | Major sigma-70 factor signature | IPR000943 | RNA polymerase sigma-70 | 292 | 307 | 5.9E-28 |
| Pfam | PF00140 | Sigma-70 factor, region 1.2 | IPR009042 | RNA polymerase sigma-70 region 1.2 | 61 | 94 | 2.7E-13 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.