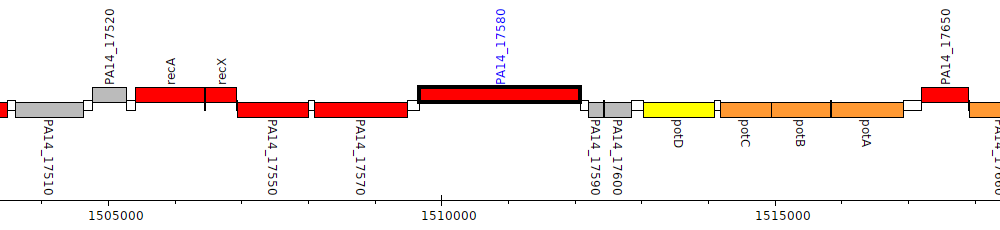
Pseudomonas aeruginosa UCBPP-PA14, PA14_17580
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016832 | aldehyde-lyase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF017245
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0005975 | carbohydrate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF017245
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PIRSF | PIRSF017245 | Phosphoketolase | IPR005593 | Xylulose 5-phosphate/Fructose 6-phosphate phosphoketolase | 26 | 801 | 0.0 |
| SUPERFAMILY | SSF52518 | Thiamin diphosphate-binding fold (THDP-binding) | IPR029061 | Thiamin diphosphate-binding fold | 420 | 620 | 2.74E-7 |
| Pfam | PF09364 | XFP N-terminal domain | IPR018970 | Xylulose 5-phosphate/Fructose 6-phosphate phosphoketolase, N-terminal | 66 | 307 | 1.1E-13 |
| Gene3D | G3DSA:3.40.50.970 | - | - | - | 425 | 634 | 3.7E-58 |
| Pfam | PF03894 | D-xylulose 5-phosphate/D-fructose 6-phosphate phosphoketolase | IPR005593 | Xylulose 5-phosphate/Fructose 6-phosphate phosphoketolase | 434 | 603 | 6.1E-43 |
| PANTHER | PTHR31273 | PHOSPHOKETOLASE-RELATED | IPR005593 | Xylulose 5-phosphate/Fructose 6-phosphate phosphoketolase | 23 | 777 | 0.0 |
| Gene3D | G3DSA:3.40.50.970 | - | - | - | 54 | 398 | 7.6E-61 |
| SUPERFAMILY | SSF52518 | Thiamin diphosphate-binding fold (THDP-binding) | IPR029061 | Thiamin diphosphate-binding fold | 81 | 387 | 1.53E-20 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.