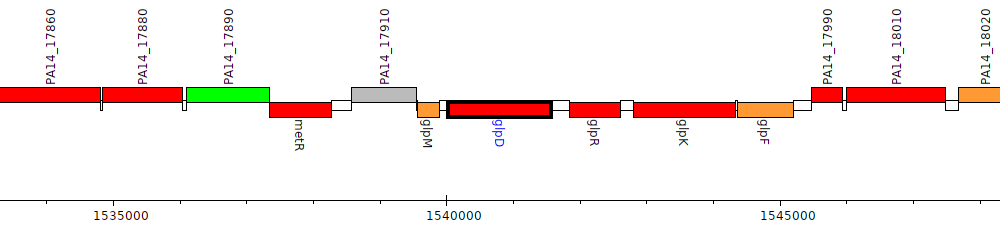
Pseudomonas aeruginosa UCBPP-PA14, PA14_17930 (glpD)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006072 | glycerol-3-phosphate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR11985
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004368 | glycerol-3-phosphate dehydrogenase (quinone) activity |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR11985
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0009331 | glycerol-3-phosphate dehydrogenase complex |
Inferred from Sequence Model
Term mapped from: InterPro:PTHR11985
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF01266
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | glycerol-3-phosphate shuttle | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | glycerophosphodiester degradation | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pau00564 | Glycerophospholipid metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | glycerol degradation I | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.30.9.10 | - | - | - | 110 | 330 | 0.0 |
| PRINTS | PR01001 | FAD-dependent glycerol-3-phosphate dehydrogenase family signature | IPR000447 | FAD-dependent glycerol-3-phosphate dehydrogenase | 44 | 56 | 2.3E-29 |
| Pfam | PF16901 | C-terminal domain of alpha-glycerophosphate oxidase | IPR031656 | Alpha-glycerophosphate oxidase, C-terminal | 391 | 486 | 1.7E-13 |
| PRINTS | PR01001 | FAD-dependent glycerol-3-phosphate dehydrogenase family signature | IPR000447 | FAD-dependent glycerol-3-phosphate dehydrogenase | 357 | 369 | 2.3E-29 |
| PANTHER | PTHR11985 | GLYCEROL-3-PHOSPHATE DEHYDROGENASE | IPR000447 | FAD-dependent glycerol-3-phosphate dehydrogenase | 8 | 486 | 7.3E-109 |
| PRINTS | PR01001 | FAD-dependent glycerol-3-phosphate dehydrogenase family signature | IPR000447 | FAD-dependent glycerol-3-phosphate dehydrogenase | 28 | 38 | 2.3E-29 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 15 | 398 | 0.0 |
| PRINTS | PR01001 | FAD-dependent glycerol-3-phosphate dehydrogenase family signature | IPR000447 | FAD-dependent glycerol-3-phosphate dehydrogenase | 89 | 101 | 2.3E-29 |
| Pfam | PF01266 | FAD dependent oxidoreductase | IPR006076 | FAD dependent oxidoreductase | 16 | 369 | 2.4E-49 |
| PRINTS | PR01001 | FAD-dependent glycerol-3-phosphate dehydrogenase family signature | IPR000447 | FAD-dependent glycerol-3-phosphate dehydrogenase | 15 | 27 | 2.3E-29 |
| Gene3D | G3DSA:6.10.250.1890 | - | - | - | 472 | 509 | 7.8E-11 |
| Gene3D | G3DSA:1.10.8.870 | - | IPR038299 | Alpha-glycerophosphate oxidase, C-terminal domain superfamily | 402 | 471 | 3.8E-28 |
| SUPERFAMILY | SSF51905 | FAD/NAD(P)-binding domain | IPR036188 | FAD/NAD(P)-binding domain superfamily | 15 | 306 | 7.93E-33 |
| FunFam | G3DSA:1.10.8.870:FF:000002 | Glycerol-3-phosphate dehydrogenase | - | - | 401 | 471 | 1.4E-29 |
| PRINTS | PR01001 | FAD-dependent glycerol-3-phosphate dehydrogenase family signature | IPR000447 | FAD-dependent glycerol-3-phosphate dehydrogenase | 323 | 329 | 2.3E-29 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.