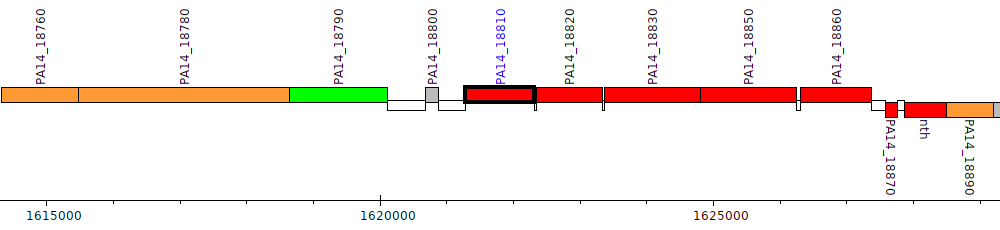
Pseudomonas aeruginosa UCBPP-PA14, PA14_18810
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0004553 | hydrolase activity, hydrolyzing O-glycosyl compounds |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:1.10.1240.20
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0042597 | periplasmic space |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:1.10.1240.20
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SMART | SM01236 | Haem_oxygenas_2_2 | - | - | 187 | 289 | 2.7E-20 |
| SUPERFAMILY | SSF48613 | Heme oxygenase-like | IPR016084 | Haem oxygenase-like, multi-helical | 120 | 334 | 7.79E-31 |
| PANTHER | PTHR40279 | PQQC-LIKE PROTEIN | IPR039068 | Pyrroloquinoline-quinone synthase-like | 84 | 344 | 7.4E-106 |
| Gene3D | G3DSA:1.10.1240.20 | Lytic transglycosylase, superhelical linker domain | IPR037061 | Lytic transglycosylase, superhelical linker domain superfamily | 21 | 106 | 2.4E-24 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 1 | 25 | - |
| Pfam | PF14518 | Iron-containing redox enzyme | - | - | 154 | 321 | 5.5E-14 |
| Pfam | PF14515 | Haem-oxygenase-associated N-terminal helices | IPR029474 | Haem-oxygenase-associated, N-terminal helices | 20 | 106 | 1.6E-43 |
| Gene3D | G3DSA:1.20.910.10 | - | IPR016084 | Haem oxygenase-like, multi-helical | 118 | 341 | 6.9E-49 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.