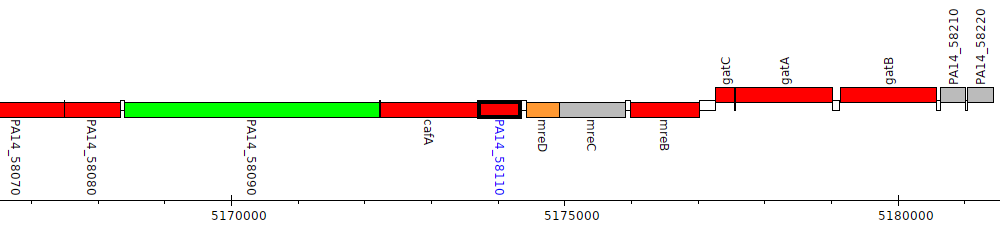
Pseudomonas aeruginosa UCBPP-PA14, PA14_58110
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0047429 | nucleoside-triphosphate diphosphatase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF006305
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.90.950.10 | - | IPR029001 | Inosine triphosphate pyrophosphatase-like | 1 | 191 | 4.1E-66 |
| PANTHER | PTHR43213 | BIFUNCTIONAL DTTP/UTP PYROPHOSPHATASE/METHYLTRANSFERASE PROTEIN-RELATED | IPR003697 | Nucleoside triphosphate pyrophosphatase Maf-like protein | 4 | 188 | 3.0E-44 |
| Pfam | PF02545 | Maf-like protein | IPR003697 | Nucleoside triphosphate pyrophosphatase Maf-like protein | 4 | 187 | 1.4E-56 |
| Hamap | MF_00528 | dTTP/UTP pyrophosphatase. | IPR003697 | Nucleoside triphosphate pyrophosphatase Maf-like protein | 3 | 190 | 29.338871 |
| NCBIfam | TIGR00172 | JCVI: Maf family nucleotide pyrophosphatase | IPR003697 | Nucleoside triphosphate pyrophosphatase Maf-like protein | 2 | 186 | 1.2E-53 |
| SUPERFAMILY | SSF52972 | ITPase-like | IPR029001 | Inosine triphosphate pyrophosphatase-like | 2 | 187 | 2.55E-59 |
| PIRSF | PIRSF006305 | Maf | IPR003697 | Nucleoside triphosphate pyrophosphatase Maf-like protein | 1 | 194 | 8.1E-50 |
| CDD | cd00555 | Maf | IPR003697 | Nucleoside triphosphate pyrophosphatase Maf-like protein | 4 | 185 | 3.44378E-87 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.