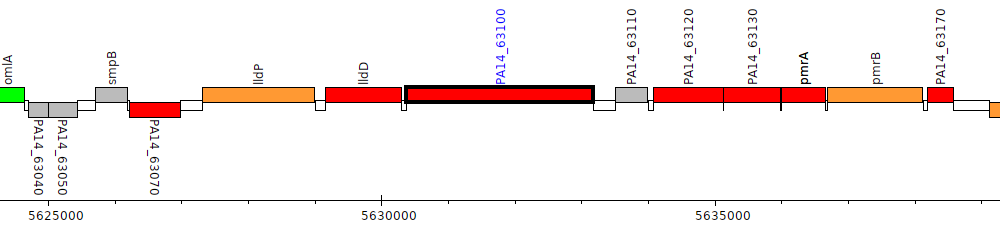
Pseudomonas aeruginosa UCBPP-PA14, PA14_63100
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0051536 | iron-sulfur cluster binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:1.10.1060.10
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0050660 | flavin adenine dinucleotide binding |
Inferred from Sequence Model
Term mapped from: InterPro:SSF55103
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003824 | catalytic activity |
Inferred from Sequence Model
Term mapped from: InterPro:SSF55103
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:1.10.45.10 | - | IPR016171 | Vanillyl-alcohol oxidase, C-terminal subdomain 2 | 476 | 515 | 1.5E-7 |
| Gene3D | G3DSA:3.30.70.2740 | - | - | - | 391 | 475 | 7.8E-11 |
| FunFam | G3DSA:1.10.1060.10:FF:000019 | Oxidoreductase/iron-sulfur cluster-binding protein | - | - | 518 | 626 | 6.9E-56 |
| SUPERFAMILY | SSF56176 | FAD-binding/transporter-associated domain-like | IPR036318 | FAD-binding, type PCMH-like superfamily | 7 | 264 | 9.0E-58 |
| SUPERFAMILY | SSF55103 | FAD-linked oxidases, C-terminal domain | IPR016164 | FAD-linked oxidase-like, C-terminal | 269 | 521 | 6.05E-58 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 671 | 694 | - |
| Gene3D | G3DSA:3.30.70.2190 | - | - | - | 267 | 383 | 3.5E-6 |
| Gene3D | G3DSA:3.30.43.10 | - | IPR016167 | FAD-binding, type PCMH, subdomain 1 | 2 | 90 | 7.8E-8 |
| Gene3D | G3DSA:3.30.465.10 | - | IPR016169 | FAD-binding, type PCMH, subdomain 2 | 96 | 265 | 5.2E-24 |
| FunFam | G3DSA:3.30.70.2740:FF:000006 | NAD-independent D-lactate dehydrogenase | - | - | 391 | 475 | 1.4E-44 |
| Pfam | PF01565 | FAD binding domain | IPR006094 | FAD linked oxidase, N-terminal | 41 | 175 | 5.1E-33 |
| FunFam | G3DSA:3.30.43.10:FF:000018 | D-lactate dehydrogenase (Dld) | - | - | 1 | 91 | 2.2E-40 |
| Pfam | PF02913 | FAD linked oxidases, C-terminal domain | IPR004113 | FAD-binding oxidoreductase/transferase, type 4, C-terminal | 267 | 514 | 6.4E-63 |
| Pfam | PF02754 | Cysteine-rich domain | IPR004017 | Cysteine-rich domain | 833 | 893 | 0.012 |
| FunFam | G3DSA:1.10.45.10:FF:000001 | D-lactate dehydrogenase mitochondrial | - | - | 476 | 516 | 8.1E-11 |
| SUPERFAMILY | SSF46548 | alpha-helical ferredoxin | - | - | 492 | 620 | 1.36E-20 |
| PANTHER | PTHR11748 | D-LACTATE DEHYDROGENASE | - | - | 9 | 522 | 3.4E-103 |
| Gene3D | G3DSA:1.10.1060.10 | - | IPR009051 | Alpha-helical ferredoxin | 518 | 626 | 2.2E-15 |
| Pfam | PF13183 | 4Fe-4S dicluster domain | IPR017896 | 4Fe-4S ferredoxin-type, iron-sulphur binding domain | 537 | 608 | 3.5E-9 |
| Pfam | PF02754 | Cysteine-rich domain | IPR004017 | Cysteine-rich domain | 704 | 795 | 0.0063 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.