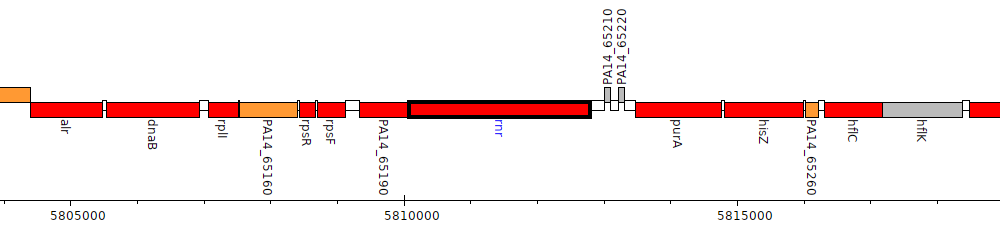
Pseudomonas aeruginosa UCBPP-PA14, PA14_65200 (rnr)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0003676 | nucleic acid binding |
Inferred from Sequence Model
Term mapped from: InterPro:SM00357
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0016070 | RNA metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR00358
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004518 | nuclease activity |
Inferred from Sequence Model
Term mapped from: InterPro:TIGR02063
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0004540 | ribonuclease activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF00773
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0003723 | RNA binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00773
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pau03018 | RNA degradation | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF50249 | Nucleic acid-binding proteins | IPR012340 | Nucleic acid-binding, OB-fold | 142 | 239 | 6.43E-18 |
| Gene3D | G3DSA:2.40.50.140 | - | IPR012340 | Nucleic acid-binding, OB-fold | 78 | 152 | 2.0E-18 |
| SMART | SM00955 | RNB_2 | IPR001900 | Ribonuclease II/R | 264 | 598 | 0.0 |
| SMART | SM00357 | csp_8 | IPR011129 | Cold shock domain | 90 | 149 | 5.1E-12 |
| Pfam | PF00575 | S1 RNA binding domain | IPR003029 | S1 domain | 659 | 740 | 6.2E-11 |
| Pfam | PF08461 | Ribonuclease R winged-helix domain | IPR013668 | Ribonuclease R winged-helix domain | 27 | 81 | 5.0E-8 |
| Pfam | PF17876 | Cold shock domain | IPR040476 | RNase II/RNase R, cold shock domain | 169 | 242 | 9.5E-23 |
| Pfam | PF08206 | Ribonuclease B OB domain | IPR013223 | Ribonuclease B, N-terminal OB domain | 91 | 148 | 5.5E-22 |
| Hamap | MF_01895 | Ribonuclease R [rnr]. | IPR011805 | Ribonuclease R | 42 | 742 | 57.126091 |
| CDD | cd04471 | S1_RNase_R | - | - | 660 | 742 | 2.57169E-43 |
| FunFam | G3DSA:2.40.50.140:FF:000362 | Ribonuclease R | - | - | 662 | 768 | 4.8E-53 |
| PANTHER | PTHR23355 | RIBONUCLEASE | - | - | 142 | 752 | 0.0 |
| SUPERFAMILY | SSF50249 | Nucleic acid-binding proteins | IPR012340 | Nucleic acid-binding, OB-fold | 661 | 741 | 2.46E-20 |
| FunFam | G3DSA:2.40.50.140:FF:000213 | Ribonuclease R | - | - | 72 | 150 | 3.7E-29 |
| Gene3D | G3DSA:2.40.50.140 | - | IPR012340 | Nucleic acid-binding, OB-fold | 662 | 771 | 1.0E-20 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 750 | 781 | - |
| Coils | Coil | Coil | - | - | 626 | 646 | - |
| Pfam | PF00773 | RNB domain | IPR001900 | Ribonuclease II/R | 264 | 597 | 4.7E-105 |
| SUPERFAMILY | SSF50249 | Nucleic acid-binding proteins | IPR012340 | Nucleic acid-binding, OB-fold | 224 | 658 | 0.0 |
| SMART | SM00316 | S1_6 | IPR022967 | RNA-binding domain, S1 | 659 | 742 | 3.9E-11 |
| MobiDBLite | mobidb-lite | consensus disorder prediction | - | - | 840 | 907 | - |
| NCBIfam | TIGR02063 | JCVI: ribonuclease R | IPR011805 | Ribonuclease R | 23 | 742 | 0.0 |
| SUPERFAMILY | SSF50249 | Nucleic acid-binding proteins | IPR012340 | Nucleic acid-binding, OB-fold | 71 | 151 | 1.52E-17 |
| NCBIfam | TIGR00358 | JCVI: VacB/RNase II family 3'-5' exoribonuclease | IPR004476 | Ribonuclease II/ribonuclease R | 79 | 741 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.