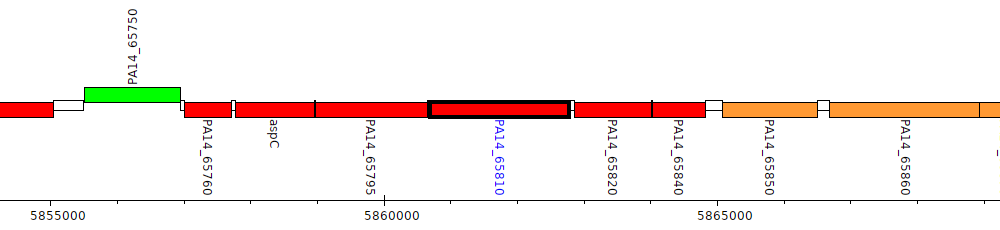
Pseudomonas aeruginosa UCBPP-PA14, PA14_65810
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0005524 | ATP binding |
Inferred from Sequence Model
Term mapped from: InterPro:G3DSA:3.30.1490.20
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | reductive TCA cycle II | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | TCA cycle V (2-oxoglutarate:ferredoxin oxidoreductase) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | TCA cycle II (plants and fungi) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | anaerobic energy metabolism (invertebrates, mitochondrial) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | pyruvate fermentation to acetate V | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | partial TCA cycle (obligate autotrophs) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | methylaspartate cycle | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | pyruvate fermentation to acetate VI | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Gene3D | G3DSA:3.40.50.261 | - | IPR016102 | Succinyl-CoA synthetase-like | 291 | 462 | 1.7E-24 |
| SUPERFAMILY | SSF52210 | Succinyl-CoA synthetase domains | IPR016102 | Succinyl-CoA synthetase-like | 129 | 286 | 7.06E-32 |
| Gene3D | G3DSA:3.30.470.20 | - | - | - | 481 | 691 | 1.3E-57 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 3 | 126 | 5.5E-25 |
| Pfam | PF13549 | ATP-grasp domain | - | - | 478 | 693 | 6.5E-52 |
| Gene3D | G3DSA:3.30.1490.20 | - | IPR013815 | ATP-grasp fold, subdomain 1 | 499 | 575 | 1.3E-57 |
| SUPERFAMILY | SSF56059 | Glutathione synthetase ATP-binding domain-like | - | - | 481 | 692 | 5.5E-30 |
| PANTHER | PTHR42793 | COA BINDING DOMAIN CONTAINING PROTEIN | - | - | 1 | 463 | 1.3E-114 |
| SMART | SM00881 | CoA_binding_2 | IPR003781 | CoA-binding | 9 | 99 | 1.5E-18 |
| Pfam | PF13380 | CoA binding domain | IPR003781 | CoA-binding | 25 | 130 | 3.5E-9 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 6 | 127 | 9.46E-23 |
| Pfam | PF13607 | Succinyl-CoA ligase like flavodoxin domain | IPR032875 | Succinyl-CoA synthetase-like, flavodoxin domain | 152 | 285 | 2.2E-33 |
| Gene3D | G3DSA:3.40.50.261 | - | IPR016102 | Succinyl-CoA synthetase-like | 127 | 286 | 1.2E-31 |
| SUPERFAMILY | SSF52210 | Succinyl-CoA synthetase domains | IPR016102 | Succinyl-CoA synthetase-like | 292 | 454 | 8.61E-22 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.