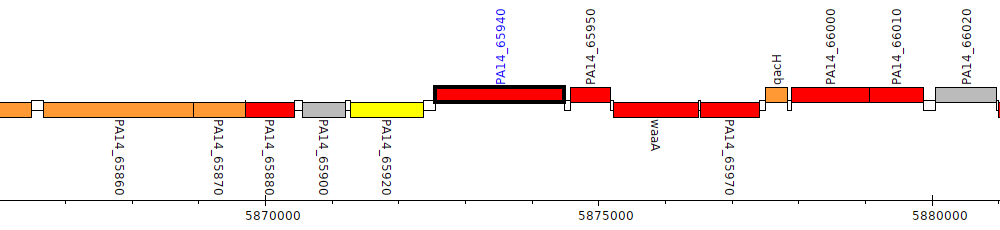
Pseudomonas aeruginosa UCBPP-PA14, PA14_65940
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0016491 | oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF07992
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0010181 | FMN binding |
Inferred from Sequence Model
Term mapped from: InterPro:PF00724
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF51395 | FMN-linked oxidoreductases | - | - | 1 | 368 | 1.08E-94 |
| Pfam | PF07992 | Pyridine nucleotide-disulphide oxidoreductase | IPR023753 | FAD/NAD(P)-binding domain | 385 | 614 | 2.8E-10 |
| SUPERFAMILY | SSF51905 | FAD/NAD(P)-binding domain | IPR036188 | FAD/NAD(P)-binding domain superfamily | 363 | 648 | 3.5E-30 |
| PANTHER | PTHR42917 | 2,4-DIENOYL-COA REDUCTASE | - | - | 1 | 534 | 0.0 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 387 | 406 | 1.8E-7 |
| PRINTS | PR00368 | FAD-dependent pyridine nucleotide reductase signature | - | - | 469 | 487 | 1.8E-7 |
| Gene3D | G3DSA:3.50.50.60 | - | IPR036188 | FAD/NAD(P)-binding domain superfamily | 481 | 605 | 4.6E-69 |
| Gene3D | G3DSA:3.20.20.70 | Aldolase class I | IPR013785 | Aldolase-type TIM barrel | 1 | 380 | 7.7E-107 |
| Pfam | PF00724 | NADH:flavin oxidoreductase / NADH oxidase family | IPR001155 | NADH:flavin oxidoreductase/NADH oxidase, N-terminal | 6 | 339 | 2.1E-59 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 386 | 641 | 4.6E-69 |
| CDD | cd04734 | OYE_like_3_FMN | - | - | 6 | 348 | 0.0 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.