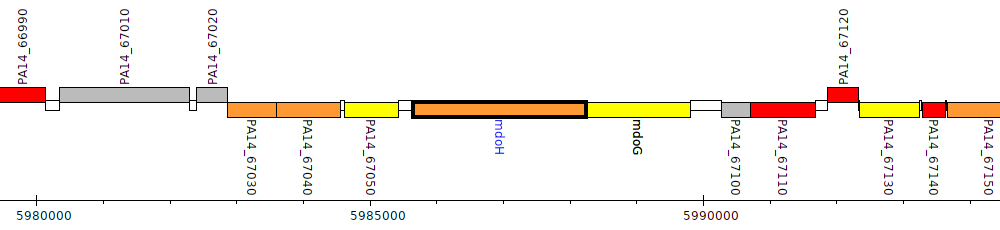
Pseudomonas aeruginosa UCBPP-PA14, PA14_67065 (mdoH)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0009405 | pathogenesis | Inferred from Mutant Phenotype | ECO:0000015 mutant phenotype evidence |
9989496 | Reviewed by curator |
| Biological Process | GO:1900727 | osmoregulated periplasmic glucan biosynthetic process |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA5077
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
17906125 | |
| Biological Process | GO:0044010 | single-species biofilm formation |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA5077
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
17906125 | |
| Molecular Function | GO:0016758 | transferase activity, transferring hexosyl groups |
Inferred from Sequence Model
Term mapped from: InterPro:MF_01072
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF00535 | Glycosyl transferase family 2 | IPR001173 | Glycosyltransferase 2-like | 249 | 431 | 1.1E-23 |
| PANTHER | PTHR43867 | CELLULOSE SYNTHASE CATALYTIC SUBUNIT A [UDP-FORMING] | - | - | 67 | 823 | 1.3E-98 |
| CDD | cd04191 | Glucan_BSP_MdoH | - | - | 247 | 500 | 0.0 |
| SUPERFAMILY | SSF53448 | Nucleotide-diphospho-sugar transferases | IPR029044 | Nucleotide-diphospho-sugar transferases | 247 | 533 | 8.06E-29 |
| Hamap | MF_01072 | Glucans biosynthesis glucosyltransferase H [mdoH]. | IPR023725 | Glucan biosynthesis glucosyltransferase H | 72 | 748 | 52.273266 |
| FunFam | G3DSA:3.90.550.10:FF:000047 | Glucans biosynthesis glucosyltransferase H | - | - | 240 | 471 | 0.0 |
| Gene3D | G3DSA:3.90.550.10 | Spore Coat Polysaccharide Biosynthesis Protein SpsA; Chain A | IPR029044 | Nucleotide-diphospho-sugar transferases | 244 | 471 | 2.3E-15 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.