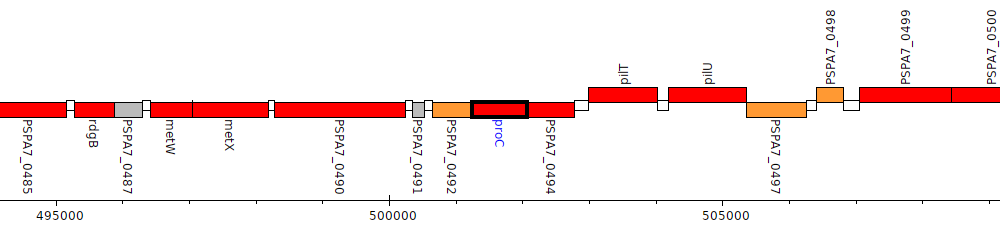
Pseudomonas aeruginosa PA7, PSPA7_0493 (proC)
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0006561 | proline biosynthetic process |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA0393
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
8479442 | |
| Molecular Function | GO:0004735 | pyrroline-5-carboxylate reductase activity |
Inferred from Sequence or Structural Similarity
Term mapped from: PseudoCAP:PA0393
|
ECO:0000249 sequence similarity evidence used in automatic assertion |
8479442 | |
| Molecular Function | GO:0004735 | pyrroline-5-carboxylate reductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000193
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006561 | proline biosynthetic process |
Inferred from Sequence Model
Term mapped from: InterPro:PIRSF000193
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| MetaCyc | L-proline biosynthesis II (from arginine) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| KEGG | pap01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pap01230 | Biosynthesis of amino acids | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pap01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pap00330 | Arginine and proline metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pap01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| MetaCyc | L-proline biosynthesis III | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
||
| MetaCyc | L-ornithine degradation II (Stickland reaction) | InterPro 5.36-75.0 |
ECO:0000259
match to InterPro signature evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| Pfam | PF03807 | NADP oxidoreductase coenzyme F420-dependent | IPR028939 | Pyrroline-5-carboxylate reductase, catalytic, N-terminal | 5 | 99 | 2.7E-23 |
| FunFam | G3DSA:1.10.3730.10:FF:000001 | Pyrroline-5-carboxylate reductase | - | - | 166 | 270 | 1.6E-34 |
| Hamap | MF_01925 | Pyrroline-5-carboxylate reductase [proC]. | IPR000304 | Pyrroline-5-carboxylate reductase | 4 | 267 | 27.789879 |
| SUPERFAMILY | SSF48179 | 6-phosphogluconate dehydrogenase C-terminal domain-like | IPR008927 | 6-phosphogluconate dehydrogenase-like, C-terminal domain superfamily | 162 | 270 | 1.87E-35 |
| Pfam | PF14748 | Pyrroline-5-carboxylate reductase dimerisation | IPR029036 | Pyrroline-5-carboxylate reductase, dimerisation domain | 163 | 267 | 1.3E-37 |
| Gene3D | G3DSA:3.40.50.720 | - | - | - | 3 | 164 | 3.0E-53 |
| SUPERFAMILY | SSF51735 | NAD(P)-binding Rossmann-fold domains | IPR036291 | NAD(P)-binding domain superfamily | 5 | 160 | 1.16E-38 |
| PANTHER | PTHR11645 | PYRROLINE-5-CARBOXYLATE REDUCTASE | - | - | 5 | 270 | 1.6E-73 |
| NCBIfam | TIGR00112 | JCVI: pyrroline-5-carboxylate reductase | IPR000304 | Pyrroline-5-carboxylate reductase | 6 | 267 | 1.3E-78 |
| FunFam | G3DSA:3.40.50.720:FF:000105 | Pyrroline-5-carboxylate reductase | - | - | 1 | 164 | 5.6E-60 |
| PIRSF | PIRSF000193 | P5CR | IPR000304 | Pyrroline-5-carboxylate reductase | 3 | 270 | 1.7E-83 |
| Gene3D | G3DSA:1.10.3730.10 | - | - | - | 165 | 271 | 3.5E-40 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.