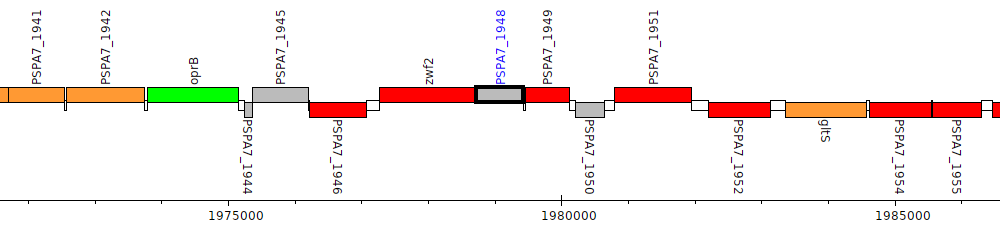
Pseudomonas aeruginosa PA7, PSPA7_1948
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Biological Process | GO:0005975 | carbohydrate metabolic process |
Inferred from Sequence Model
Term mapped from: InterPro:cd01400
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006098 | pentose-phosphate shunt |
Inferred from Sequence Model
Term mapped from: InterPro:cd01400
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Molecular Function | GO:0017057 | 6-phosphogluconolactonase activity |
Inferred from Sequence Model
Term mapped from: InterPro:cd01400
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
Pathways
| Database | Xref | Pathway | Version | Evidence | PMID |
|---|---|---|---|---|---|
| KEGG | pap01120 | Microbial metabolism in diverse environments | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pap00030 | Pentose phosphate pathway | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pap01110 | Biosynthesis of secondary metabolites | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pap01200 | Carbon metabolism | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pap01130 | Biosynthesis of antibiotics | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
|
| KEGG | pap01100 | Metabolic pathways | 81.0+/01-23, Jan 17 |
ECO:0000249
sequence similarity evidence used in automatic assertion |
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| SUPERFAMILY | SSF100950 | NagB/RpiA/CoA transferase-like | IPR037171 | NagB/RpiA transferase-like | 16 | 232 | 7.67E-54 |
| Pfam | PF01182 | Glucosamine-6-phosphate isomerases/6-phosphogluconolactonase | IPR006148 | Glucosamine/galactosamine-6-phosphate isomerase | 19 | 236 | 1.8E-50 |
| CDD | cd01400 | 6PGL | IPR005900 | 6-phosphogluconolactonase, DevB-type | 21 | 238 | 2.29304E-74 |
| PANTHER | PTHR11054 | 6-PHOSPHOGLUCONOLACTONASE | IPR039104 | 6-Phosphogluconolactonase | 14 | 236 | 1.7E-29 |
| Gene3D | G3DSA:3.40.50.1360 | - | - | - | 14 | 238 | 1.4E-59 |
| NCBIfam | TIGR01198 | JCVI: 6-phosphogluconolactonase | IPR005900 | 6-phosphogluconolactonase, DevB-type | 17 | 233 | 1.8E-71 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.