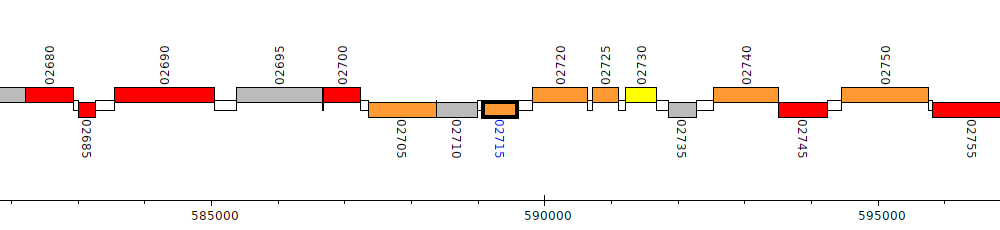
Pseudomonas aeruginosa DK2, PADK2_02715
Cytoplasmic
Cytoplasmic Membrane
Periplasmic
Outer Membrane
Extracellular
Unknown
Gene Ontology
| Ontology | Accession | Term | GO Evidence | Evidence Ontology (ECO) Code | Reference | Comments |
|---|---|---|---|---|---|---|
| Molecular Function | GO:0015035 | protein disulfide oxidoreductase activity |
Inferred from Sequence Model
Term mapped from: InterPro:PF02600
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Biological Process | GO:0006457 | protein folding |
Inferred from Sequence Model
Term mapped from: InterPro:PF02600
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
||
| Cellular Component | GO:0016020 | membrane |
Inferred from Sequence Model
Term mapped from: InterPro:PF02600
|
ECO:0000259 match to InterPro signature evidence used in automatic assertion |
Functional Classifications Manually Assigned by PseudoCAP
|
|
Functional Predictions from Interpro
| Analysis | Accession | Description | Interpro Accession | Interpro Description | Amino Acid Start | Amino Acid Stop | E-value |
|---|---|---|---|---|---|---|---|
| PANTHER | PTHR36570 | DISULFIDE BOND FORMATION PROTEIN B | - | - | 1 | 163 | 3.5E-48 |
| SUPERFAMILY | SSF158442 | DsbB-like | IPR023380 | DsbB-like superfamily | 13 | 159 | 1.05E-32 |
| Hamap | MF_00286 | Disulfide bond formation protein B [dsbB]. | IPR022920 | Disulphide bond formation protein DsbB | 6 | 168 | 18.331043 |
| Gene3D | G3DSA:1.20.1550.10 | - | IPR023380 | DsbB-like superfamily | 1 | 167 | 9.3E-37 |
| Pfam | PF02600 | Disulfide bond formation protein DsbB | IPR003752 | Disulphide bond formation protein DsbB/BdbC | 11 | 159 | 1.5E-33 |
Search for additional functional domains at the NCBI CDD database website. Go to this protein's amino acid sequence and follow the link.